Extracellular DNase activity was studied in 73 strains of Cryptococcus neoformans and 12 strains of Cryptococcus gattii. DNase activity was measured by DNase agar clearance with and without Methyl Green. All strains tested showed extracellular DNase activity and no significant difference was found betweenC. neoformans and C. gattii strains. DNase production was higher in strains from clinical origin (average radius of 6.2mm) than among environmental strains (average radius of 2.9mm). The extracellular enzyme may be detected by DNA substrate PAGE assays and its molecular weight was estimated at 31kD. These results suggest that extracellular DNase could be considered as a virulence factor involved in C. neoformans–C. gattii species complex pathogenicity.
Se ha estudiado la actividad extracelular de la enzima ADNasa en 73 cepas de Cryptococcus neoformans y 12 cepas de C. gattii. Para medir dicha actividad se utilizó la técnica de aparición de halos claros en agar DNasa con y sin verde metilo. Todas las cepas analizadas produjeron DNasa extracelular sin observar diferencias significativas entre C. neoformans y C. gattii. La producción de DNasa fue mayor en los aislamientos clínicos (valor medio del halo: 6,2mm) que en los recogidos de medio ambiente (valor medio del halo: 2,9mm). Mediante ensayo en gel PAGE con ADN como sustrato, se pudo estimar el peso molecular del enzima en 31kD. Estos resultados permiten apoyar la hipótesis de que la ADNasa extracelular podría ser un factor de patogenicidad implicado en la virulencia en el complejo de especies C. neoformans-C. gattii.
Cryptococcus neoformans and Cryptococcus gattii are fungal pathogens that cause cryptococcosis. Until recently, both microorganisms were considered varieties of the same species, i.e., C. neoformans var. neoformans and C. neoformans var. gattii, however now they are recognized as separated species19, due to their differences in morphology, ecology, physiology, genetics and serology.1,18 The two species also show different pathogenic behaviours. C. neoformans primarily affects immunocompromised patients, like those who suffer AIDS,14 causing pneumonia and meningitis. C. gattii is considered a primary pathogen causing cryptococcosis in immunocompetent individuals. For both pathogens, infection is acquired from the environment, presumably through the respiratory route22, and both species produce a panoply of extracellular enzymes27 like proteinases,4,8 urease,20,32 phenoloxidase28 and phospholipases.9,10,33,34
Extracellular DNase production by Cryptococcus was described several years ago6, and its possible role in pathogenesis has been previously discussed5 - DNase from C. neoformans, however, is best known as the “usual suspect” for the difficulties encountered in generating high molecular weight DNA from Cryptococcus for molecular studies.5 The characteristics of that enzyme in these yeasts have not been studied thus far. In this study we approached the question as to whether pathogenic isolates of C. neoformans and C. gattii are associated with extracellular DNase activity. By testing numerous isolates of the two species in DNase agar we found that both species produce extracellular DNase, but clinical isolates of C. neoformans produce significantly more extracellular DNase than environmental isolates. These results indicate that Cryptococcus DNase may contribute to in virulence by degrading host DNA.
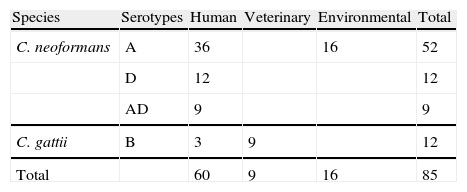
Material and methodsStrainsC. neoformans and C. gattii strains used in this study were isolated from different environmental and clinical sources (Table 1). Most of the strains used were previously described.2,11–13,16 Seven Uruguayan clinical isolates (five C. neoformans serotype A, one serotype D and one C. gattii serotype B), were donated by Dr. Nora Fernández. The strains are maintained in the stock culture collection of the Laboratory of Mycology at the University Miguel Hernández (CCA—Colección Crytococcus Alicante, Spain). All were serotyped and tested for other phenotypical features like urease, phenoloxidase and capsule production, ability to grow at 37°C and assimilation of different carbohydrates (auxonograme). The isolates were recovered in liquid YED prior to the inoculation on DNase agar plates. Saccharomyces cerevisiae CLA024 (bakery commercial strain) was used as a negative control.
DNase agar clearanceThe method used for detecting extracellular DNase production by yeasts was similar to the procedure used for detecting DNase production by staphylococci. Plates of DNase Test Agar (Oxoid) supplemented with methyl green were prepared.29 Five microliters from the liquid cultures described above were inoculated in the plates and incubated at 30°C for 7 days. After that time, clear zones were observed around the colonies. The strains showed differences in the colony size. Hence the halo was measured by the radius from the edge of the colony. To increase the clearance of the halos, the plates were flooded with 1N hydrochloric acid. After 1min, pictures of the plates were taken for halo measuring. Assays were repeated at least twice for each strain and the average radius was determined. Data were analyzed with Student's t-test and significance was defined as P<0.001.
DNA substrate PAGECCA197 strain, which showed the highest DNase production, was selected for the characterization of the enzyme. After its culture on liquid YED the supernatant was recovered and the protein was obtained by precipitation with ammonium sulfate (AS). Three fractions were obtained: 50%, 75% and 100% of saturation with AS. Precipitates were resuspended in 500μl of PBS and dialyzed against the same buffer overnight. DNase activity in the samples was determined by the development of clearance halo in plates of DNase Test Agar after adding 20μl of the different fractions. DNase activity was observed in the 75% and 100% fractions. Non-reducing sodium dodecyl sulfate polyacrilamide gel electroforesis (SDS-PAGE) was used to visualize DNase activity in DNA containing gels as described previously.3 Twenty-five microliters of those fractions positive for DNase were loaded into each lane. Samples were not boiled to avoid destruction of activity. Sigma DNaseI was used as a control. Following electrophoresis the gels were incubated in a magnesium buffer to allow renaturation and DNase activity to take place. After ethidium bromide staining, DNase activity could be detected as dark areas that appeared under UV exposition.
ResultsAll the strains tested showed DNase activity. No activity was detected with the S. cerevisiae strain used as negative control.
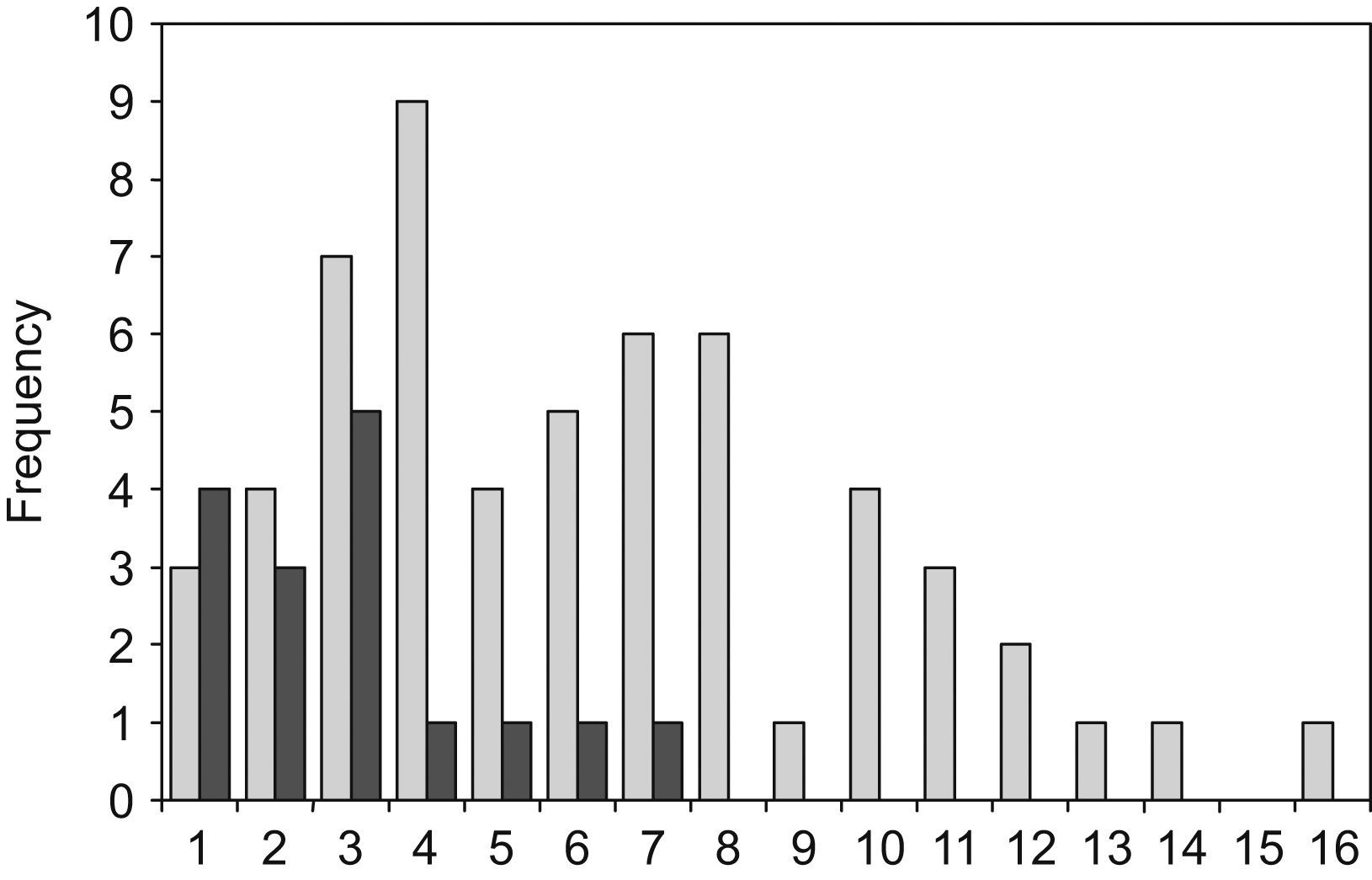
The radius of the clearance halo from the edge of the colony varied from 1 to 16mm among the different strains. CCA197 and CCA209 strains, both clinical isolates of C. neoformans, produced the largest clearance zones. The smallest halos were observed in four environmental, two veterinarian and four clinical isolates.
C. neoformans environmental isolates were a very homogeneous group, which produced the smallest halos (16 strains, with an average radius of 2.9mm; σ=1.7). The C. neoformans clinical isolates were a more heterogeneous group and produced wider halos (57 strains with an average radius of 6.2mm; σ=3.5). As shown in Fig. 1 the difference between the environmental and clinical strains is statistically significant (t=3.63). Only those isolates obtained from human patients showed a radius of over 8mm. This large halo was found in 13 isolates of C. neoformans. There was a tendency for a bimodal distribution of the radius size in clinical isolates, although no statistical significance was found with the number of samples used in this study. The halos produced by C. gattii group, that included the 9 isolates from veterinarian origin, had an intermediate size between the values noted above (12 strains with an average radius of 4.2mm; σ=2.3). When comparing the average DNase halos between C. neoformans and C. gattii species, no significant difference was found. Additionally, no relationship was found between the DNase activity and the serotypes of the strains.
DNA substrate PAGE analysis showed a major dark area of DNase activity at an apparent molecular weight of 31kD (not shown). Commercial DNase I showed an apparent molecular weight of 20kD, which is smaller than its real weight (around 31kD). This difference can be explained because samples were not boiled and the electrophoresis was performed in non-reducing conditions. Thus, the true molecular weight of cryptococcal DNase could be somewhat higher.
DiscussionMembers of the C. neoformans–C. gattii complex have been considered both primary and secondary pathogens. They can produce primary disease in apparently normal immunocompetent individuals, which is not rare in C. gattii infections, and more commonly cause secondary disease in severely immunosupressed hosts. In addition, asymptomatic cryptococcal infections are frequent and most do not cause diseases because the host's immune system which controls the infection.27
Virulence phenotypes for these species are well characterized. The three most important phenotypes are capsule production, melanin formation, and the ability of the strain to grow well at 37°C, and these have been shown to be under genetic control5,27 . Nonetheless other mechanisms like the production of several extracellular enzymes, have been considered as important virulence factors.5,27,28 Phospholipase aids in tissue invasion and in the escape from the phagosome by breaking down host cell membranes.17,25 Extracellular protease exerts a similar activity and its production has also been described in pathogenic strains.8,33,34 Urease is another extracellular enzyme that is considered a virulent factor facilitating the invasion of the central nervous system.26,32 Phenoloxidase is involved in the previously mentioned melanin synthesis and consequently, it protects the cell from oxidative attack.28
Although DNase was described several years ago as an extracellular enzyme produced by Cryptococcus,5,6 no further studies have been carried out thus far regarding the role of the enzyme, if any, in virulence. Extracellular DNase is considered as a virulence factor in bacterial pathogenesis.23,24,30,35 Extracellular DNase activity has been described as required for the normal progression of Group A Streptococcus infection by enhancing evasion of the innate immune system, specifically by avoiding the killing caused by neutrophils.31 In cryptococcosis, neutrophils have been observed in the inflammatory response to the infection and found in close association with the infected tissue.15,21 Therefore, it is possible that cryptococcal DNase acts in a way similar to estreptococal DNase.
The results described here show that the enzyme is produced by environmental and clinical isolates but clinical isolates produce significantly higher levels of this enzyme. Considering that many other non-pathogenic fungi produce extracellular DNases7,36 and that C. neoformans–C. gattii are also free-living organisms, it appears that the yeasts, in their environmental niche, may have developed adaptive tools to protect themselves from soil amoebas, and other biotic factors, as well as to obtain nutrients from the environment. These protective mechanisms enhance survival but produce disease in the mammalian host.27 Therefore, production of DNase and other extracellular enzymes could play a role in wild life, but it seems that the yeasts enhance some of these abilities during their parasitic life in human environment. This hypothesis would need to be demonstrated by in vivo and in vitro studies. Nevertheless, we can conclude that members of the C. neoformans–C. gattii complex produce an extracellular DNase of an apparent molecular weight of 31kD, which may be a virulence factor when the yeast parasitizes the human body.
This work was supported by a grant from Fundación Navaro-Tripodi, Alicante, Spain. We are grateful to Dr. Nora Fernandez for the Uruguayan Cryptococcus strains, Rocio Valera and Concepción Nuñez for their excellent technical assistance, and María Abad for her English language corrections.