The virulence of isolates among different Candida species causing candidemia may play a role in the prognosis of the patients. Furthermore, the potential relationship between genotype and virulence is still unclear and need to be further studied.
AimsWe aim to assess the relationship between genotype and virulence in Candida species using a Galleria mellonella larvae infection model.
MethodsOne hundred and ninety-four isolates from 68 clusters (Candida albicans, 114/41; Candida parapsilosis, 74/24; Candida tropicalis, 6/3) were compared against the same number of each species singleton genotypes in terms of survival of G. mellonella larvae.
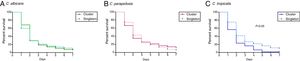
ResultsThe median of survival and the IQR ranges of clusters and singleton were as follows: C. albicans (2 days, IQR 1.5–2 vs. 2 days, IQR 1–2.25), C. parapsilosis (2 days, IQR 1.5–2.6 vs. 2 days, IQR 2–3.3), and C. tropicalis (1 day, IQR 1–3.5 vs. 2 days, IQR 2–3.5; p<0.05). High intra-cluster variability in terms of median of survival was found regardless the species.
ConclusionsNo relationship between genotype and virulence in Candida was observed with the G. mellonella model.
La virulencia de cepas de diferentes especies de Candida causantes de candidemia puede jugar un papel en el pronóstico de los pacientes, y su estudio en el modelo de infección en Galleria mellonella puede ser útil para entender su contribución general a la infección. Además, la potencial relación entre genotipo y virulencia requiere de más estudios.
ObjetivosSe evaluó la relación entre genotipo y virulencia en especies de Candida mediante el modelo de infección de larvas de G. mellonella.
MétodosSe estudió la supervivencia de las larvas infectadas con 194 aislados incluidos en 68 clusters (Candida albicans, 114/41; Candida parapsilosis, 74/24; Candida tropicalis, 6/3) y con el mismo número de aislados con genotipos únicos por especie.
ResultadosLa mediana de supervivencia y los rangos intercuartílicos (IQR) de clusters y genotipos únicos se muestra a continuación: C. albicans (2 días, IQR: 1,5-2 vs. 2 días, IQR: 1-2,25), C. parapsilosis (2 días, IQR: 1,5-2,6 vs. 2 días, IQR: 2-3,3), y C. tropicalis (un día, IQR: 1-3,5 vs. 2 días, IQR: 2-3,5; p<0,05). Encontramos una importante variabilidad en la mediana de supervivencia entre cepas del mismo cluster, independientemente de la especie analizada.
ConclusionesNo se encontró relación entre el genotipo y la virulencia entre los aislados de Candida evaluados mediante el modelo de infección de G. mellonella.
Candidemia is primarily caused by Candida albicans, Candida parapsilosis, Candida glabrata, and Candida tropicalis.5 The virulence of Candida isolates, such as adherence and biofilm formation, the ability to evade the host's immune system, and the production of tissue-damaging hydrolytic enzymes (proteases, phospholipases, and haemolysins), may play a role in the prognosis of infected patients.10,11Galleria mellonella has become a useful model of infection for testing different pathogens as larvae are easy to handle, are cheap, and their innate immune system resembles that of mammals.14 However, the relationship between genotype and virulence is mostly unknown.1,8,9,12,13 We previously showed intra- and inter-species differences in terms of virulence among Candida strains using a G. mellonella infection model,8 and that clusters present greater biofilm production than singleton isolates.3 Here we evaluated the potential relationship between genotype and virulence in Candida by studying Candida cluster and singleton genotypes using a G. mellonella model.
We studied 68 clusters of Candida species causing candidemia in patients admitted to 16 tertiary hospitals in Spain, Denmark, Italy and Brazil during 2014–2015. Isolates had been genotyped previously using species-specific microsatellite markers.3,4 A cluster was defined as an identical genotype infecting at least two patients. Comparisons of 194 isolates from clusters (C. albicans, n=114 isolates from 41 clusters; C. parapsilosis, n=74 isolates from 24 clusters; C. tropicalis, n=6 isolates from 3 clusters) and other 194 isolates with singleton genotype (one isolate per genotype) from patients with candidemia admitted to Gregorio Marañón hospital, were carried out. Virulence was established by assessing the survival in the G. mellonella larvae infection model as previously reported.8 Briefly, each Candida inoculum was adjusted to 5×107cells/ml (C. albicans and C. tropicalis) or 2×108–5×108cells/ml (C. parapsilosis) with a Neubauer hematocytometer. Each experimental group containing 10 larvae of G. mellonella (Bichosa, Salceda de Caselas, Spain) with similar size and weigh were infected with 10μl of each cell suspension. Ten larvae PBS-inoculated and 10 non-injected larvae were used as controls. Larvae were incubated at 37°C, and death was determined visually for seven days. Isolate survival curves were generated using Kaplan–Meier method. Median survival was calculated and log-rank (Mantel–Cox test) was applied to examine the differences between the survival curves (Graph Pad Prism statistical 5.02 software, GraphPad, La Jolla, USA). The interquartile range (IQR) of median survival was calculated per species, and for clusters and singletons, by pooling the isolates. Isolates were further classified as highly virulent (median survival<percentil25), low virulent (median survival>percentile75) and moderately virulent (in between) according to the pooled analysis of clusters and singleton isolates. This study was approved by the Ethics Committee of Hospital Gregorio Marañón (CEIC-A1; study no. 201/18).
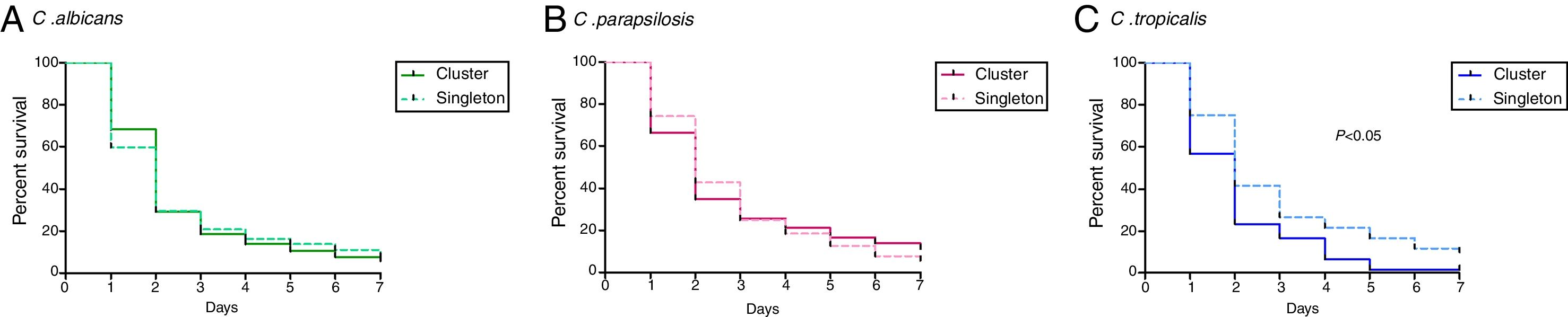
PBS control larvae and those non-injected were alive the seven days post-infection. Overall, the pooled median survival times of larvae infected by Candida species were the following: C. albicans (2 days, IQR 1–2), C. tropicalis (2 days, IQR 1.13–3.13), and C. parapsilosis (2 days, IQR 2–3). Differences in median survival between larvae infected by cluster isolates and singletons did not reach statistical significance except for C. tropicalis (Fig. 1; p<0.05). Mean of survival and IQR ranges of clusters and singleton were as follows: C. albicans (2 days, IQR 1.5–2 vs. 2 days, IQR 1–2.25; p>0.05), C. parapsilosis (2 days, IQR 1.5–2.6 vs. 2 days, IQR 2–3.3; p>0.05), and C. tropicalis (1 day, IQR 1–3.5 vs. 2 days, IQR 2–3.5; p<0.05). High intra-cluster variability in terms of mean survival was found regardless the species. Based on median survival data, clusters vs. singleton isolates were classified as highly virulent [C. albicans (0% vs. 0%), C. parapsilosis (28.4% vs. 10.8%), and C. tropicalis (50% vs. 0%)], moderately virulent [C. albicans (76% vs. 75.4%), C. parapsilosis (48.6% vs. 64.9%), and C. tropicalis (16.7% vs. 86.3%)], and low virulent [C. albicans (3.7% vs. 24.6%), C. parapsilosis (23% vs. 24.3%), and C. tropicalis (33.3% vs. 16.7%).
This study supports previous observations of higher mortality associated to C. albicans and C. tropicalis in comparison to C. parapsilosis in both G. mellonella models and clinical observations.5,8 Moreover, we aimed to decipher potential intra-species differences in terms of virulence between clusters and singletons. Differences reached statistical significance only for C. tropicalis, although the low number of isolates due to the lower number of cases of candidemia caused by this species (n=12) did not allow us to reach solid conclusions.4,5 We carried out a species-specific classification of isolates based on G. mellonella mortality. Subtle differences were observed, despite the fact that we found significantly higher percentages of highly virulent isolates of C. parapsilosis and C. tropicalis in clusters compared with singletons. These observations suggest that genotypes and mortality in G. mellonella are unrelated. In previous studies, contradictory conclusions were reported when the genotype was compared with virulence factors not related to G. mellonella mortality. Some authors found positive correlations,6,9,12,15 and others did no find such relationship.1,7,13 One of the most studied virulence factors is biofilm formation. C. albicans biofilm producing isolates are more aggressive than non-biofilm producer isolates in the G. mellonella larvae model,2 although we were unable to initially support these observations in a previous study.8 Subsequently, we included genotyping in the equation and observed a tendency of C. albicans clusters to form more biofilm than singletons.3 However, the huge intra-cluster variability found in terms of biofilm formation,3 along with the variability in terms of mortality in G. mellonella reported here, suggest notable isolate-to-isolate differences rather than a genotype–phenotype virulence relationship. Our study is subject to certain limitations. First, we have a heterogeneous number of isolates per species. Second, since whole genome sequencing of these isolates has not yet been performed, the variability among isolates in clusters can be a consequence of the lack of discriminatory potential for the microsatellites. Finally, we were unable to include any clinical data of patients and make a correlation between our in vitro findings and patient outcome.
In conclusion, no relationship between genotype and virulence in G. mellonella infection model was found for the three studied Candida species, suggesting that epigenetic factors are important determinants of virulence.
Author contributionsJudith Díaz-García: methodology; formal analysis; writing – original draft preparation. Maiken C. Arendrup and Rafael Cantón: data collection; resources (samples); writing –review & editing. Julio García-Rodríguez, Elia Gómez García de la Pedrosa, Gabriella Parisi, Javier Pemán, Brunella Posteraro, Maurizio Sanguinetti, Daniel Archimedes Da Matta, Arnaldo L. Colombo, Patricia Muñoz, Carlos Sánchez-Carrillo: data collection; resources (samples) and writing. Jesús Guinea and Pilar Escribano: conceptualization; project administration; data collection; supervision; validation; visualization; writing – original draft preparation and review & editing.
FundingThis study was supported by grants PI16/01012 and CP15/00115 from Fondo de Investigación Sanitaria (FIS, Instituto de Salud Carlos III, Plan Nacional de I+D+I 2013–2016). The study was co-funded by the European Regional Development Fund (FEDER) ‘A way of making Europe.’ In Brazil, this work was supported by a grant 2017/02203-7 from Fundação de Amparo a Pesquisa-São Paulo (FAPESP).
PE (MS15/00115) is a recipient of a Miguel Servet contract supported by FIS. JG is a steady researcher contracted by Fundación para Investigación Sanitaria del Hospital Gregorio Marañón. JD is supported by a predoctoral grant by FIS (FI19/00021).
Conflict of interestJG has received funds for participating at educational activities organized on behalf of Astellas, Gilead, Pfizer, MSD, Scynexis, and Biotoscana-United Medical; he has also received research funds from FIS, Gilead, Scynexis, Cidara, Basilea Ltd, and F2G outside the submitted work. ALC received educational grants from Biotoscana-United Medical, MSD, Pfizer and a research grant from Astellas and Pfizer. RC has received funds for participating at educational activities organized by MSD and Pfizer. MCA reports personal fees from Astellas, Basilea, Gilead, MSD, Pfizer, T2Biosystems, and Novartis, and others from Amplyx, Astellas, Basilea, Gilead, T2Biosystems, F2G, Novabiotics and Scynexis outside the submitted work. The rest of the authors have no conflicts of interests.
The authors are grateful to Dainora Jaloveckas (cienciatraducida.com) for the editing assistance.