To characterize one linezolid- and methicillin-resistant Staphylococcus aureus (MRSA) isolate recovered from a nasal sample of a pig farmer patient.
MethodsThe detection of linezolid resistance mechanisms was performed by PCR and sequencing. The antimicrobial resistance and virulence profile was investigated, and the molecular typing was performed by molecular techniques. The transference of cfr gene was assessed by conjugation experiments and its genetic environment was investigated by specific PCRs.
ResultsThe linezolid-resistant MRSA isolate was typed as t011-ST398/CC398-SCCmecV-agrI and carried the cfr gene. The isolate was multidrug-resistant but lacked the virulence genes studied. The cfr gene was co-located with the fexA gene on a Tn558 variant and was successfully transferred by conjugation.
ConclusionWe report the first description of LA-MRSA-CC398 carrying the cfr gene in Spain. This finding highlights the importance of surveillance programmes to determine the presence and spread of the cfr gene in the livestock and clinical settings.
Caracterizar una cepa Staphylococcus aureus resistente a meticilina (SARM) y a linezolid aislada de una muestra nasal de un paciente granjero de cerdos.
MétodosMediante PCR y secuenciación se investigaron los mecanismos de resistencia a linezolid. Se determinó el perfil de resistencia a antimicrobianos y el perfil de virulencia, y se llevó a cabo el tipado molecular mediante diferentes técnicas moleculares. Se estudió la transferibilidad del gen cfr por conjugación y su entorno genético mediante PCR específicas.
ResultadosEl aislado fue tipado como t011-ST398/CC398-SCCmecV-agrI y portaba el gen cfr. Presentó un feno/genotipo de multirresistencia, pero carecía de los genes de virulencia estudiados. Se detectó el gen cfr junto con fexA en una variante del Tn558 y se transfirió mediante conjugación.
ConclusiónSe describe el primer aislado SARM-CC398 cfr-positivo en España. Esto destaca la importancia de implementar programas de vigilancia para determinar su presencia y dispersión en el ámbito clínico y ganadero.
Staphylococcus aureus clonal complex CC398 is the most common genetic lineage of livestock-associated (LA) methicillin-resistant S. aureus (MRSA) in Europe. Pigs are the main host of MRSA-CC398, however, its detection in humans is increasing in recent years either as colonizer or as causative agent of infection.1
Besides methicillin resistance, MRSA-CC398 usually exhibits a multiresistance phenotype which limits the therapeutic options. In this context, linezolid is an important antimicrobial agent for the treatment of serious infections caused by Gram-positive bacteria, including MRSA. Point mutations in the domain V of the 23S rRNA, as well as amino acid changes in the ribosomal proteins L3, L4, and L22 are the most common mechanisms of linezolid resistance in staphylococci.2 Moreover, linezolid resistance mediated by acquired resistance genes is of concern due to they are often located on mobile genetic elements, which can move between bacteria by horizontal gene transfer. To date, three transferable linezolid resistance genes have been described in staphylococcal species: cfr, optrA, and poxtA.3–5 Furthermore, these genes also confer reduced susceptibility to other antimicrobial agents, such as phenicols.2
In this study, we report, as far as we know, the first description of MRSA-CC398 harbouring the cfr gene in Spain, recovered from a nasal sample of a patient with professional contact with pigs.
MethodsA pig farmer was hospitalized in the Intensive Care Unit (ICU) of the Hospital Universitari Arnau de Vilanova (Lleida, Spain) due to a non-infectious disease. Active surveillance for multiresistant bacteria was performed upon ICU admission, and MRSA were isolated from nasal, pharyngeal, and bronchoalveolar lavage samples, that were considered as colonization.
The susceptibility to penicillin, oxacillin, erythromycin, clindamycin, gentamicin, tobramycin, amikacin, minocycline, tetracycline, ciprofloxacin, levofloxacin, vancomycin, teicoplanin, fosfomycin, mupirocin, rifampicin, fusidic acid, daptomycin, and trimethoprim-sulfamethoxazole was determined by commercialized broth microdilution method (MicroScan, Beckman Coulter, Brea, USA) and to cefoxitin by disc diffusion.6 In addition, the minimum inhibitory concentration (MIC) to linezolid was determined by E-test® (bioMérieux, Durham, USA), and to chloramphenicol6 and florfenicol7 by broth microdilution using S. aureus ATCC® 29213 as quality control. All MRSA isolates recovered showed identical antimicrobial resistance phenotype and the isolate recovered from the nasal sample, named X1761, was further characterized.
The presence of the linezolid resistance genes cfr, cfr(B), optrA, and poxtA was investigated by PCR (Supplementary Table S1). Moreover, mutations and amino acid changes in the deduced sequence of 23S rRNA and in the genes encoding the ribosomal proteins L3 (rplC), L4 (rplD), and L22 (rplV) were determined by PCR and amplicon sequencing (Supplementary Table S1). The obtained sequences were compared with those of linezolid-susceptible S. aureus subsp. aureus ATCC® 25923 (GenBank accession number CP009361) using the EMBOSS Needle software for nucleotide or amino acid (BLOSUM 62 cost matrix) alignments.
Based on the antimicrobial resistance phenotype, the presence of the following antimicrobial resistance genes was studied by PCR: mecA, mecB, mecC, blaZ, lnu(A/A′), lnu(B), lsa(B), vga(A), tet(K), tet(L), tet(M), ant(4′)-Ia, aac(6′)-Ie-aph(2″)-Ia, fexA, fexB, catpC194, catpC221, and catpC223 (Supplementary Table S1). Moreover, amino acid changes in the genes encoding the GyrA and GrlA proteins were investigated by PCR and amplicon sequencing (Supplementary Table S1) and compared with those of the fluoroquinolone-susceptible strain S. aureus subsp. aureus ATCC® 25923 (GenBank accession number CP009361). The presence of the virulence factors Panton-Valentine leukocidin (lukS/F-PV), toxic shock syndrome toxin-1(tst), and exfoliative toxins (eta, etb, and etd) was determined by PCR (Supplementary Table S1).
The MRSA isolate was subjected to spa-typing by PCR and amplicon sequencing, and the obtained sequence was analyzed using Ridom Staph-Type software (Ridom GmbH, Münster, Germany). Moreover, the isolate was characterized by agr-typing, multilocus sequence typing, and staphylococcal cassette chromosome mec (SCCmec) typing (Supplementary Table S1).
The transferability of the cfr gene was evaluated by the filter-mating method. A rifampicin-resistant mutant of S. aureus ATCC® 29213 was used as recipient strain, and the selection of the transconjugants was performed on BHI agar containing linezolid (4mg/L) and rifampicin (50mg/L). The transconjugants were identified by spa-typing. The antimicrobial resistance phenotype and genotype was determined by broth microdilution and by PCR, respectively (Supplementary Table S1).
The genetic environment of the cfr gene was determined by PCR mapping and amplicon sequencing. The primers were designed based on the structure of the cfr-carrying plasmids pSCFS3 and pSCFS7 (GenBank accession number AM086211 and FR675942) (Supplementary Table S2).
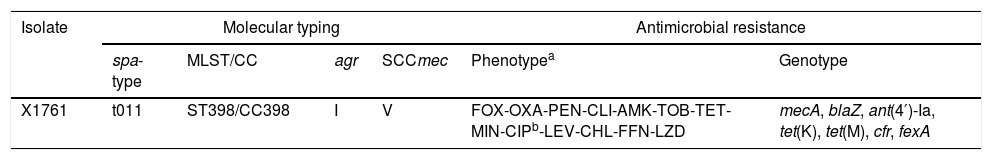
Results and discussionThe MRSA isolate was resistant to thirteen of the twenty-three antimicrobial agents tested, including linezolid (MIC 8mg/L), but remained susceptible to vancomycin and daptomycin. It carried the mecA, blaZ, ant(4′)-Ia, tet(K), tet(M), cfr, and fexA resistance genes and showed the amino acid changes S80F and S84L in GrlA and GyrA proteins, respectively, related to fluoroquinolone resistance (Table 1). The cfr gene, which encodes a rRNA methyltransferase, mediates combined resistance to phenicols, lincosamides, oxazolidinones, pleuromutilins, and streptogramins A antimicrobial agents. This multiresistance gene was originally detected in the plasmid pSCFS1 from S. sciuri,3 and, thereafter, in different Gram-positive and Gram-negative bacteria of diverse origins. Although the use of linezolid is not approved in food-producing animals, the cfr gene is spread among staphylococcal isolates from pigs in different countries such as China,8 which might be due to the co-selection of resistances. The linezolid-resistant MRSA isolate of this study also displayed the amino acid change V118A in the deduced sequence of rplD gene. There is no data about the relation of this change with linezolid susceptibility, but it seems to be associated with decreased susceptibility to the pleuromutillin lefamulin in S. aureus isolates.9
Molecular typing, and antimicrobial resistance phenotype and genotype of the linezolid-resistant MRSA X1761 isolate.
| Isolate | Molecular typing | Antimicrobial resistance | ||||
|---|---|---|---|---|---|---|
| spa-type | MLST/CC | agr | SCCmec | Phenotypea | Genotype | |
| X1761 | t011 | ST398/CC398 | I | V | FOX-OXA-PEN-CLI-AMK-TOB-TET-MIN-CIPb-LEV-CHL-FFN-LZD | mecA, blaZ, ant(4′)-Ia, tet(K), tet(M), cfr, fexA |
The X1761 isolate was typed as t011-ST398/CC398-SCCmecV-agrI. MRSA-CC398 isolates carrying the cfr gene have been described sporadically in different countries recovered primarily from livestock, but also from humans.10,11 However, as far as we know, this is the first detection of MRSA CC398-cfr in Spain. A recent study conducted among Spanish hospitals, demonstrated the correlation between the pig density of the region and the presence of MRSA-CC398 in human infections at hospital level.1 The hospital in which this strain was obtained is located in the Spanish region with the highest pig density in the year 2019 (370 pigs/km2); moreover, the patient was a pig farm worker, which is known to be a risk factor for human colonization with MRSA-CC398.12
The isolate did not harbour any of the virulence determinants studied, which is commonly observed among LA-MRSA CC398 isolates in comparison with human-adapted lineages.13 The isolate lacked the IEC genes, which suggests an animal origin and, therefore, an animal to human transmission.
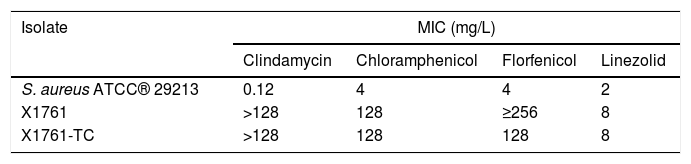
The cfr gene was successfully transferred by conjugation into S. aureus ATCC® 29213 together with the fexA gene. The transconjugants showed reduced susceptibility to clindamycin, chloramphenicol, florfenicol, and linezolid (Table 2), which is in accordance with the antimicrobial resistance genotype detected. The PCR mapping allowed us to identify the cfr and fexA genes as part of the Tn558 variant described on the plasmid pSCFS3 of S. aureus (GenBank accession number AM086211),14 where the cfr gene is flanked by an IS21-558 element. The nucleotide alignment of the cfr genetic environment of the S. aureus X1761 isolate and the Tn558 variant of the plasmid pSCFS3 revealed the presence of six nucleotide exchanges in the non-coding region upstream the IS21-558 element. Although the cfr gene has been reported integrated in the chromosomal DNA, it has been more frequently detected on plasmids and, in many cases, with other resistance genes, including fexA.15 Kehrenberg and Schwarz14 described that the partial deletion of the transposase genes tnpA and tnpB in the Tn558 variant, due to the integration of the IS21-558 element, resulted in a transposition-deficient structure. Hence, the transference of the cfr gene by conjugation indicates its possible location on a plasmid, what would facilitate its dissemination.
In conclusion, the presence of transferable linezolid resistance genes in MRSA recovered from humans represents a public health concern, since it is considered a key antimicrobial agent in human medicine. MRSA-CC398 is a genetic lineage of the human-animal interface that frequently colonize pigs and is disseminated to in-contact humans. Consequently, the detection of a cfr-positive MRSA CC398 strain colonizing the nose of a pig farmer patient suggests that this multidrug-resistant strain might be circulating within the pig farms, what should be monitored in the future.
FundingThis work was partially supported by project SAF2016-76571-R from the Agencia Estatal de Investigación (AEI) of Spain and the Fondo Europeo de Desarrollo Regional (FEDER) of EU. Laura Ruiz-Ripa has a pre-doctoral fellowship from the Universidad de La Rioja (Spain).
Conflict of interestThe authors declare no conflict of interest.