Bordetella pertussis is the causative agent of pertussis, which mainly affects unvaccinated children, while Bordetella parapertussis causes a disease presenting clinical characteristics that are indistinguishable from whooping cough. Despite high vaccination coverage, pertussis remains a public health concern worldwide, with approximately 140000 cases reported annually. Here we determined the prevalence of B. pertussis and B. parapertussis infection among infants under one year of age by polymerase chain reaction (PCR); our aim being to identify whether the data obtained relates to the relevant sociodemographic and clinical data. The study included 86 samples of nasopharyngeal swabs from infants aged between 0–12 months, who were reported as probable cases of whooping cough by the health centers around the Ecuadorian highlands, from August 2016 to July 2017. The nasopharyngeal swabs were cultured and microbiological and molecular analyses were performed. B. pertussis was identified by PCR in 41% of the samples (30/86), more than half of which corresponded to infants aged between 0–3 months. Moreover, a statistically significant correlation (p<0.05) between the identification of bacteria in culture and the catarrhal stage of the disease was observed. The results obtained from the study highlighted the need for an active national surveillance of pertussis, in particular for laboratory testing, to provide a highly sensitive and more specific diagnosis of Bordetella infection.
Bordetella pertussis es el agente causal de la tosferina, que afecta principalmente a población pediátrica no vacunada; mientras que Bordetella parapertussis ocasiona un cuadro clínico similar indistinguible de tosferina. A pesar de la alta cobertura de vacunación, pertussis sigue siendo un problema de salud pública a nivel mundial, con aproximadamente 140.000 casos reportados anualmente. Se determinó la prevalencia de la infección por B. pertussis y B. parapertussis en niños menores de un año, por reacción en cadena de la polimerasa (PCR), y se identificó la relación entre los datos obtenidos y los datos sociodemográficos y clínicos relevantes. El estudio incluyó 86 muestras nasofaríngeas de menores entre 0 a 12 meses de edad, quienes fueron reportados como casos probables de tosferina por centros de salud de la región Sierra de Ecuador, desde agosto de 2016 hasta julio de 2017. Las muestras nasofaríngeas fueron analizadas mediante el método de cultivo, pruebas microbiológicas y moleculares. Se identificó B. pertussis por PCR en el 41% de muestras (30/86), más de la mitad de estas muestras correspondieron a niños de entre 0 y tres meses de edad. Además, se identificó una correlación estadísticamente significativa (p < 0,05) entre la detección de las bacterias mediante cultivo y la fase catarral de la enfermedad. Con base en estos resultados, es necesario mejorar el sistema de vigilancia epidemiológica del país, principalmente, en la confirmación de casos por laboratorio, para una detección altamente sensible y específica del microorganismo causante de la tosferina.
During the past years, a resurgence of pertussis cases has been observed worldwide and is considered a reemerging infectious disease by the World Health Organization (WHO)26,39,46. This disease is present in developing and developed countries, despite high rates of vaccination. It is caused by gram-negative bacteria belonging to the genus Bordetella, namely, Bordetella pertussis and Bordetella parapertussis, the latter causing a less severe form of the disease3,10.
B. pertussis is known to cause more severe disease than B. parapertussis or Bordetella holmesii, while Bordetella bronchiseptica affects immunocompromised patients more frequently36. These three species of Bordetella share some virulence factors such as adhesins, fimbriae, filamentous hemagglutinin (FHA), pertactin (PRN), tracheal colonization factor, and tracheal cytotoxin. Conversely, pertussis toxin and serum resistance protein are expressed only by B. pertussis, and the type III secretion system is expressed only by B.bronchiseptica37.
Among the virulence factors, it is possible to find adenylate cyclase toxin-hemolysin (CyaA), an adhesin that is frequently included as a component in acellular vaccines against B.pertussis44. Moreover, this factor is characterized by the presence of a hemolytic phenotype of virulent strains of B. pertussis. At present, it has been reported that the hemolytic phenotype in Bordetella is lost once other genes replace the toxin.
The infection has shown a modified pattern of transmission in areas with high coverage of pertussis vaccination, which is likely due to a change in circulating B. pertussis strains4. However, according to some authors, the resurgence of pertussis cases is either a consequence of bacterial host adaptation or due to vaccine influence3. Of the vaccines available for the prevention of the disease, the Center for Disease Control and Prevention (CDC) has recommended the use of the acellular vaccine as the primary choice for infants under one year of age, as it has been proved to be less reactogenic10.
In Ecuador, based on the expanded immunization program (EPI), the pentavalent and trivalent whole-cell inactivated vaccines (WCV) are the pertussis vaccines being currently administered. However, in accordance with the biological drug release registry, there are also acellular pertussis vaccines (ACV) being provided by the private sector. Moreover, a decreased vaccination coverage has been observed; between 2001 and 2009 vaccination coverage was 100%; however, since 2009, this rate has been gradually decreasing32.
Nowadays, health organizations aim at reducing the morbidity and mortality rates in lactating infants, who are under one year of age, as this population registers the majority of pertussis cases reported worldwide (88.7 out of 10000)41. To achieve this objective, vaccination strategies should be evaluated; vaccination and booster doses should be implemented to cover more than 90% of the population. Moreover, epidemiological surveillance systems of the disease should be improved so that an increased number of probable cases of pertussis are identified and diagnosed26. In Ecuador, the Ministry of Public Health through the SIVE-Alerta program performs a passive surveillance of the disease by notifying probable cases of B. pertussis infection. In 2015, 7 cases were reported and in 2016 there was an increase of 14 cases35. Since 2016 a rapid rise in the number of cases of B. pertussis infection has been observed; in fact, they have duplicated in comparison to the previous year, which could be suggestive of a new outbreak. Nonetheless, the confirmation of positive cases is hampered due to a lack of highly sensitive and more specific tests, such as the polymerase chain reaction (PCR)22,46. This study aimed to identify the prevalence of B. pertussis, B. parapertussis and B. holmesii in infants under one year of age, who were reported as probable cases of whooping cough. Diagnosis was confirmed by microbiological culture and PCR, for faster and more specific laboratory diagnosis.
MethodsPopulation samplingA cross-sectional study was carried out in the city of Quito, Ecuador, in coordination with the Instituto Nacional de Investigación en Salud Pública (INSPI). The Ecuadorian highland region was chosen for evaluation as it has reported the highest number of pertussis cases since 201635. The samples to be analyzed consisted of 86 nasopharyngeal swabs from children between 0 to 12 months of age, who were described as probable cases of whooping cough after visiting the different health centers in the region.
The study was limited to a type of observational study that analyzes data from a population, from the Epidemiological Surveillance System for this disease. This study involves data collected at a defined time. The study was reviewed and approved by the Human Research Ethics Committee of the Pontificia Universidad Católica del Ecuador, which categorized it as scientifically acceptable.
SamplesSamples were received either as nasopharyngeal swabs in the Regan-Lowe Semi-solid Transport Medium (calcium alginate swab, one swab per nostril) or as nasopharyngeal aspirate specimens in a sterile bottle. The samples were analyzed by the gold standard method: microbiological culture, and by endpoint PCR. The latter followed an established protocol for the amplification of B. pertussis targets IS481 and ptxA-Pr, B. parapertussis IS1001 and B. holmessi hIS1001. This study was done from August 2016 to July 2017.
Data collectionThe epidemiological chart of each sample was evaluated to retrieve relevant sociodemographic and clinical data. A database was created, including the following information for each sample: the age and gender of the patient, place of origin, type of sample, elapsed time from symptom onset to primary diagnosis, which was determined by the practitioner based on the clinical symptoms.
Microbiological cultureNasopharyngeal swabs were cultured in Regan-Lowe medium (Becton Dickinson), containing 5% cephalexin, and were incubated in a humidity chamber at 37°C for 7–10 days. After this period, the presumptive Bordetella spp. colonies were subjected to further testing to differentiate B. pertussis from B. parapertussis. Microbiological and biochemical tests were used, including Gram staining, MacConkey agar, oxidase, and catalase tests.
PCR for the detection of B. pertussis and B. parapertussisEach swab sample was suspended in 500μl of PBS, and genomic DNA was extracted using the High Pure PCR Template Preparation Kit (Roche Germany), according to the manufacturer's instructions.
To differentiate among the Bordetella species, specific primers were used for the following targets: IS481 and ptxA-Pr were used for the identification of B. pertussis, IS1001, for the identification of B. parapertussis, and the hIS1001 target for the identification of B.holmesii22,40 (Table S1). To amplify the bacterial genome, 1μl of the extracted DNA was added to 11.5μl of the master mix preparation (GoTaq®) to make up a final volume of 12.5μl. The conditions used in the thermocycler were the following: denaturation (5min at 94°C); annealing for IS481 (30s at 63.6°C); annealing for ptxA-Pr (30s at 55°C); annealing for IS1001 (30s at 60.4°C); and annealing for hIS1001 (30s at 63.6°C) extension (30s at 72°C); final extension (10min at 72°C); 25 cycles were used to amplify the genes.
Statistical data analysisDescriptive statistics were used to analyze the clinical information of the patients, and the results obtained from the microbiological cultures and PCR. The non-parametric chi-square test (χ2) was used to compare the results obtained from the cultures and PCR to establish a diagnostic value. The parameters of sensitivity, specificity, positive predictive value and negative predictive value were also analyzed.
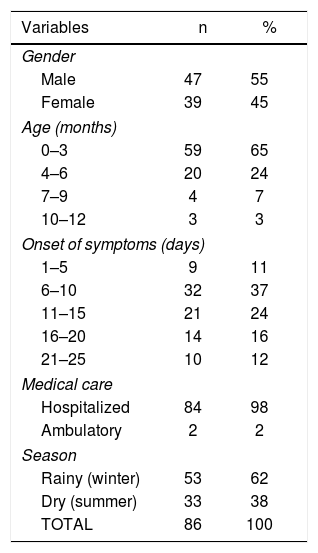
ResultsOf the 86 samples of probable cases of whooping cough, 29 (35%) belonged to patients who were hospitalized in the pediatric unit at health centers in Quito. No significant difference was observed between genders, as 55% of patients were male, and 45% were female. The study included a population ranging from 0–12 months of age, which was distributed as follows: 59 patients were between 0–3 months old; 20, between 4–6 months old; 4, between 7–9 months old, and 3 between 10–12 months old. More than half of the samples used in the study, 62 of 86 (72%), came from patients going through the first 2 weeks since symptom onset, whereas 24 (28%) were past this time frame (Table 1). From the epidemiological charts in our possession, it was not possible to verify the vaccination status or the antimicrobial treatment of the patients; therefore, these factors, if any, were not included in our analysis.
Characteristics of the study population.
| Variables | n | % |
|---|---|---|
| Gender | ||
| Male | 47 | 55 |
| Female | 39 | 45 |
| Age (months) | ||
| 0–3 | 59 | 65 |
| 4–6 | 20 | 24 |
| 7–9 | 4 | 7 |
| 10–12 | 3 | 3 |
| Onset of symptoms (days) | ||
| 1–5 | 9 | 11 |
| 6–10 | 32 | 37 |
| 11–15 | 21 | 24 |
| 16–20 | 14 | 16 |
| 21–25 | 10 | 12 |
| Medical care | ||
| Hospitalized | 84 | 98 |
| Ambulatory | 2 | 2 |
| Season | ||
| Rainy (winter) | 53 | 62 |
| Dry (summer) | 33 | 38 |
| TOTAL | 86 | 100 |
The data was retrieved from the epidemiological charts of patients reported as probable cases of whooping cough in Quito, Ecuador.
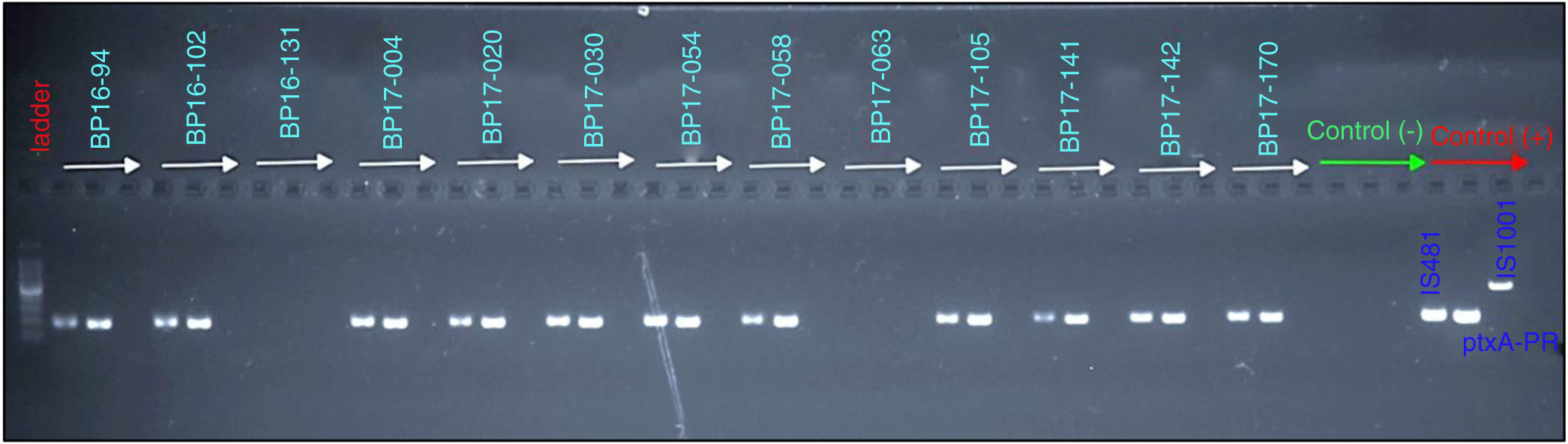
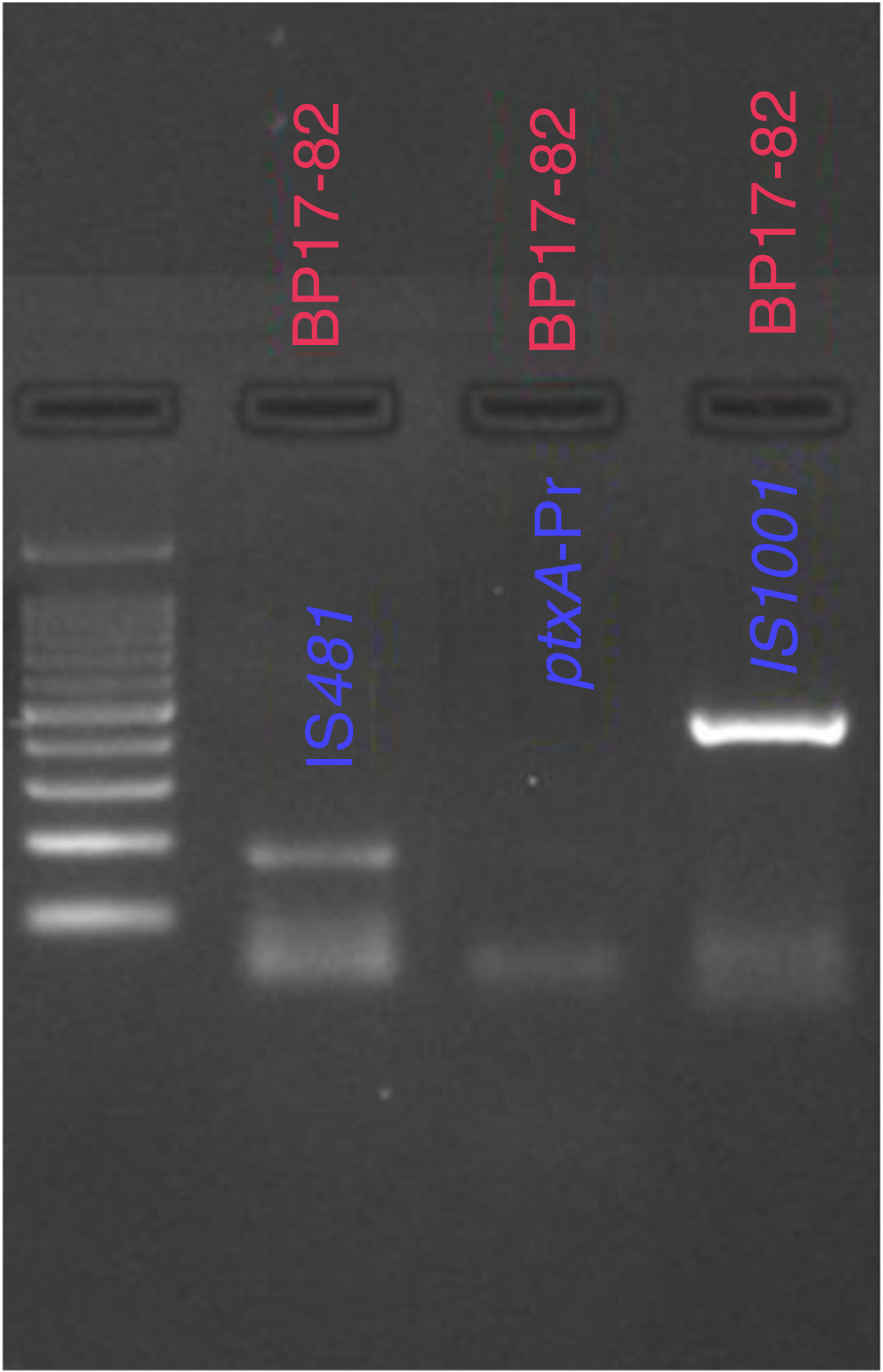
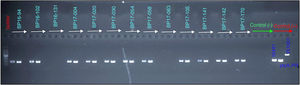
From the samples that were analyzed, 23% (20 of 86) were found to be positive for B. pertussis by culturing, whereas the results obtained from PCR showed that 41% (30 of 86) of cases were positive for IS481 and ptxA-Pr, which are specific to B. pertussis (Fig. 1). Moreover, one case was identified as a co-infection of B. pertussis and B. parapertussis, since both IS481 and IS1001 were amplified (Fig. 2), while the hIS1001 did not amplify any sample (Table S2).
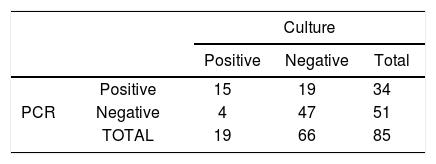
By comparing the techniques used for the detection of the bacteria, it was observed that the polymerase chain reaction showed a sensitivity of 79% and a specificity of 71%. In contrast, the gold standard method showed 100% specificity, and 23% sensitivity (20/86) (Table 2).
Lastly, a correlation between the different stages of the disease and an identification of the bacteria in culture was analyzed, and a statistically significant result (p-value <0.05) was revealed for children whose swabs were collected between 1–15 days after the onset of symptoms, otherwise known as the catarrhal stage of the disease (Table 3).
Correlation between the onset of symptoms of the population under study, and the detection of Bordetella pertussis by culture.
| Onset of symptoms (days) | Culture | ||
|---|---|---|---|
| Positive for B. pertussis | Negative for B. pertussis | Total | |
| 1–5 days | 2.30% | 8.10% | 10.50% |
| 6–10 days | 4.70% | 32.60% | 37.20% |
| 11–15 days | 11.60% | 12.80% | 24.40% |
| 16–20 days | 2.30% | 14.00% | 16.30% |
| 21–25 days | 2.30% | 9.30% | 11.60% |
| Total | 23.30% | 76.70% | 100% |
| Pearson's chi-squared test (χ2) | χ2 value: 9457; df.: 4; p-value: 0.04 | ||
Whooping cough is an endemic disease with eventual outbreaks and epidemic peaks, which manifest every 2 to 5 years13,26. During the last decade, Latin America has reported a resurgence of pertussis post-vaccination era, the annual incidence of the disease continues to rise, and the pertussis-related mortality rate in infants younger than 6 months old is estimated at 0.2%9,14,38.
In our study, the majority of positive cases of B. pertussis infections corresponded to those infants belonging to the group of 0–3 months of age, reaching a prevalence of 65%. This percentage was similar to that identified in a study carried out in Peru, in which 70% of positive cases corresponded to infants younger than 3 months old7 as one of the main risk factors for the disease at this age is incomplete vaccination2,9. Nonetheless, it is worth mentioning that neither vaccination nor natural infection confers complete, lifelong immunity to the disease42; in fact, a minimum of two doses should be administered for vaccination to be effective. For this reason, lactating infants under 4 months of age are at an increased risk of contracting the infection6,42.
Recently, there has been a resurgence of cases of pertussis caused by B. pertussis, despite good vaccination coverage1,25. This resurgence has been attributed to the introduction of the acellular vaccine (ACV)34. However, the genomic variation of pertactin (PRN) and pertussis toxin (Ptx) of B. pertussis has been described since the introduction of the whole-cell vaccine (WCV)5.
Most strains used for the production of WCV and ACV express antigens such as PtxA2 or PtxA4 and Prn1 or Prn7, whereas circulating clinical isolates have other variants, namely, PtxA1 and Prn2 or Prn335. In addition, strains of B. pertussis have been isolated from countries with high vaccination coverage, which have lost antigens such as filamentous hemagglutinin (FHA), pertactin (Prn), Ptx subunit A (PtxA), serotype 2 fimbriae (Fim2), serotype 3 fimbriae (Fim3), and tracheal colonization factor (TcfA)44. While in countries with low vaccination coverage, the emergence of new cases of pertussis has been attributed to the presence of B. pertussis strains that express a new allele for the pertussis toxin promoter, which enhances the production of the pertussis toxin (Ptx)34.
To this day, strains of B. pertussis that do not produce Prn have been isolated mainly in countries that report pertussis epidemics. In the United States, during an outbreak in 2012, it was recorded that more than 60% of isolates did not produce Prn43. Additionally; studies have shown that B. pertussis strains that do not produce Prn induce a stronger pro-inflammatory response, and increase cell death after infection23.
Given that the symptoms of the disease tend to be non-specific, clinical data are vital for the adequate diagnosis of probable cases of pertussis31. In this study, none of the epidemiological charts contained information about the vaccination status of the patient, mother or other family members, which is necessary to monitor vaccination compliance. Furthermore, there is an immunity window of vulnerability for infants under 2 months of age11,19. For this reason, the WHO considers maternal immunization as an effective strategy for the prevention of pertussis in infants, as passive immunization is reached after the transfer of maternal antibodies4,19,25.
In this study, two techniques were used for the detection of B. pertussis in our samples to evaluate the efficiency of the molecular technique PCR. It was identified that culturing yielded 23% of positive cases, whereas PCR detected 41%; this variation was expected since other studies estimated that without PCR, 9% to 26% of positive cases could be missed every year17.
The results obtained from this study have shown that the application of PCR assays for the laboratory confirmation of B. pertussis and B. parapertussis, provides increased sensitivity, as it can detect minimum amounts of bacterial DNA from a sample. Moreover, to identify and differentiate among Bordetella species, two assays were used, each designed to amplify species-specific DNA sequences20,27,45. For this reason, the CDC has recommended the use of PCR as the primary method for the detection of pertussis infection16,30.
A statistically significant correlation was observed in our study population between the catarrhal stage of the disease, which extends from day 1 to 15 after the onset of symptoms, an identification of the bacteria in culture26,29. In contrast, the use of PCR as a laboratory diagnostic tool amplified the window for the detection of the microorganism; the onset of symptoms can extend from day 1 to 30, including the catarrhal stage of the disease as well as the paroxysmal stage 2.
The different methodologies applied for the identification of pertussis allowed to extend the period in which the disease was detected, in comparison to the detection time of a test used in isolation28. Several health organizations such as the Minnesota Department of Health suggest that the optimal time for the detection of pertussis by culture should be calculated from samples collected during the first 2 weeks since the onset of symptoms33. Similarly, Leber AL in 2014 observed that cultures have a sensitivity of 12–60% if the analysis is performed at the beginning of the disease (<2 weeks)29, that is, during the catarrhal stage of the disease, when there is an increased bacterial load that favors the diagnosis by culture. Moreover, Leite et al. note that in their study the sensitivity of cultures and PCR decreased after the second week since the onset of symptoms, dropping from 70% and 80% respectively to 10% and 21%30. Infections caused by B. pertussis and B. parapertussis that cause a pertussis-like syndrome or co-infection might be undistinguishable18. For this reason, a laboratory diagnosis of the etiological agent is necessary. Recently, studies have shown that the prevalence of B. parapertussis infections has increased8; however, the pertussis vaccine provides little to no protection against B. parapertussis infection. Based on this fact, the information obtained from this research will be made available to healthcare professionals, so that decisions are made in the clinical-epidemiological setting by taking into account the frequencies identified here for the first time.
The reemergence of pertussis is a public health concern that poses a significant challenge to existing health policies. This emphasizes that there is an atypical clinical diagnosis, as the disease manifests itself differently depending on the age, stage of the disease, immunological status, or region where it is located; these factors, therefore, complicate the epidemiological surveillance of the disease in Ecuador.
In this study, we analyzed the effectiveness of laboratory diagnostic tools for the identification of pertussis. It was observed that the use of PCR was more sensitive for the detection of the bacteria than culturing, probably because it can detect and amplify a single copy of bacterial DNA21,24. It is therefore likely that the negative results yielded by culture might have been false-negatives, which further supports the increased sensitivity of PCR (79%) over the cultures12,15. Finally, the results from the study suggest the incorporation of PCR testing as a tool for early detection of whooping cough in hospitals and epidemiological surveillance in Ecuador. We provided a rapid and accurate diagnosis of the disease to reduce the time response for the treatment of the infection, especially for infants under three months of age, considering that clinical suspicion alone is often not enough to accurately diagnose the disease43,45,46.
Ethics approvalThis study has been reviewed and approved by the Ethics Committee of Human Research of the PUCE, considering it scientifically acceptable.
Conflict of interestThe authors declare that they have no conflicts of interest.
We thank Nicolle Ramírez Cifuentes for her English language advice. This study was supported and funded by the Pontificia Universidad Católica del Ecuador, Project code M13455.