Currently, the valorization of agroindustrial waste is of great interest. Moringa oleifera is a multipurpose tree whose softwood residues could be used as raw material for low-cost cellulase production. The aim of this study was to isolate, identify, and characterize microorganisms with cellulolytic activity in different carbon sources. We isolated and purified 42 microorganisms from M. oleifera biomass. Fungi presenting the largest hydrolytic halos in carboxymethylcellulose as a substrate were molecularly identified as Penicillium funiculosum (FG1), Fusarium verticillioides (FG3) and Cladosporium cladosporioides (FC2). The ability of these fungal strains to break down cellulose was assessed in a submerged fermentation using either amorphous CMC or crystalline form (Avicel). P. funiculosum and C. cladosporioides displayed similar endoglucanase (606U/l) and exoglucanase (205U/l) activities in the Avicel-containing medium, whereas F. verticillioides showed the highest level of β-glucosidase activity (664U/l) in the carboxymethylcellulose medium. In addition, the effect of three culture media (A, B, and C) on cellulase production was evaluated in P. funiculosum using moringa straw as a carbon source. The results showed a volumetric productivity improvement of cellulases that was 2.77-, 8.26-, and 2.30-fold higher for endoglucanase, exoglucanase and β-glucosidase, respectively when medium C containing moringa straw was used as a carbon source. The enzymatic extracts produced by these fungi have biotechnological potential especially for second-generation bioethanol production (2G) from moringa straw. This is the first report on the use of M. oleifera biomass to induce the production of various cellulases in P. funiculosum.
Actualmente, la valorización de los residuos agroindustriales es de gran interés. En este trabajo se emplearon residuos de madera blanda de Moringa oleifera para la producción de celulasas de bajo costo. El objetivo fue aislar, identificar y caracterizar microorganismos con actividad celulolítica en diferentes fuentes de carbono. A partir de la biomasa de M. oleifera, se aislaron e identificaron 42 microorganismos productores de celulasas. Los hongos que presentaron los mayores halos de hidrólisis en carboximetilcelulosa como sustrato fueron identificados molecularmente como Penicilliumfuniculosum (FG1), Fusariumverticillioides (FG3) y Cladosporiumcladosporioides (FC2). Mediante fermentación sumergida, se evaluó la capacidad de estas cepas en la producción de celulasas utilizando celulosa cristalina (Avicel) y amorfa (CMC) como fuentes de carbono. P. funiculosum y C. cladosporioides presentaron las mayores actividades de endoglucanasa (606 U/l) y exoglucanasa (205 U/l) en medio Avicel, mientras que F. verticillioides mostró la mayor actividad de β-glucosidasa (664 U/l) en medio CMC. Además, se evaluó el efecto de tres medios de cultivo (A, B y C) sobre la producción de celulasas en P. funiculosum empleando residuos de moringa como fuente de carbono. Los resultados mostraron que en el medio C, la productividad volumétrica de celulasas se incrementó en 2,77; 8,26 y 2,30 veces para las actividades de endoglucanasa, exoglucanasa y β-glucosidasa, respectivamente. Los extractos enzimáticos producidos tienen gran potencial para su utilización biotecnológica, especialmente en la sacarificación de residuos de moringa y la producción de bioetanol de segunda generación. Este es el primer estudio del uso de la biomasa de M. oleifera para inducir la producción de diversas celulasas en P. funiculosum.
Numerous studies have examined the possibility of large-scale ethanol production starting from lignocellulosic residues. However, the major technological issue to address is the general absence of low-cost technology that overcomes the recalcitrance of cellulosic biomass2,3,17. Lignocellulosic biomass represents a promising option as feedstock for ethanol production and for cellulase production, which must be in accordance with the interests of each country for conveying value to cellulosic wastes, especially for those without food value12,35.
Cellulose is the main component of the plant cell wall and is assembled in different orientations throughout its structure, leading to different levels of crystallinity. Such a specialized and complicated structure renders cellulose resistant to biological and chemical attacks13. A promising strategy to overcome this impediment involves the production of cellulolytic enzymes, hydrolysis of biomass, and fermentation of resulting sugars to generate desired products. Cellulase production by different cellulolytic microorganisms is being dynamically studied to reduce the cost of breaking down the plant biomass. Complete cellulose hydrolysis is mediated by a combination of three main types of cellulases: (I) endoglucanases (EnG) that randomly break down the amorphous regions of cellulose, giving rise to oligosaccharides, diminishing the length of the linear cellulose chains and increasing the reducing sugar content; (II) exoglucanases (ExG), including cellobiohydrolases that progressively cut on the reducing or non-reducing ends of cellulose chains, liberating cellobiose or glucose; and (III) β-glucosidase (BG), which hydrolyzes cellobiose and cellodextrins, yielding glucose molecules12,43.
Both fungi and bacteria have been heavily exploited for their abilities to produce a wide variety of cellulases and hemicellulases. More emphasis has been placed on the use of fungi rather than of bacteria because fungi can produce copious amounts of enzymes, which are secreted into the medium for easy extraction and purification15.
The most common fungi cited for cellulase production is Trichoderma reesei. However, its production is dependent on costly media components and T. reesei strains generally lack sufficient BG activity. This is why it is common to combine it with other fungi such as Aspergillus spp12,24. Nevertheless, there is a great biodiversity of microorganisms that has not been explored yet; in particular, little information has been provided on the microbial ecology and occurrence of viable microflora in cellulosic biomaterial26,34.
Moringa oleifera (common name moringa) is an oilseed tree that can be considered of great interest for biofuel production (biodiesel and bioethanol). It is cultivated in different tropical and subtropical countries, such as India, Cuba, and Mexico11,18. Recently, an increased interest in investigating moringa cultivation and waste utilization has been mainly focused on obtaining biotechnological products23. During moringa oil extraction several residues are generated that can be used to produce cellulosic ethanol, as recently reported by Visser et al.38 in soybean, castor bean, jatropha, palm kernel, sunflower, and cottonseed.
The aim of the present study was to isolate, identify, and characterize cellulolytic microorganisms from moringa biomass and to evaluate the effect of different carbon sources (including synthetic cellulose and lignocellulosic biomass) on the cellulolytic activities and cellulase production of the isolated microorganisms.
Materials and methodsSample collectionIn May 2013, straw (leaves and stems) of the M. oleifera were collected. Samples were collected in three different localities of the State of Sinaloa (Culiacán, Guasave and Sinaloa de Leyva), Mexico. Subsequently, they were placed in hermetically sealed bags and transported to the laboratory, where they were stored at 4°C until further use.
Isolation, screening, and identification of strainsIsolation and screeningThe tissues were washed in sterile water, cut into small pieces of 2–5mm squares, transferred by using flame-sterilized forceps to sterile petri dishes containing sodium hypochlorite (1%) for 30–60s and washed in sterile water two or three times30. The tissue treated was milled, 1g of sample was placed in 10ml of sterile water and shaken for proper mixing at room temperature and serial dilutions up to 1×10−7 were prepared in sterile distilled water for each sample. Afterwards, 0.1ml of each dilution was plated on potato dextrose agar (PDA) for fungi isolation and Luria-Bertani (LB) agar for bacteria isolation. The plates were incubated at 30°C for 4 days. The fungal cultures were purified by the single hyphal tip method and the serial dilution plate technique was used to purify bacteria30. Isolated microorganisms were qualitatively evaluated for cellulase production after incubation at 30°C for 2 days using an agar medium containing carboxymethylcellulose® (Aldrich Chemistry, CAS: 9004-32-4, Saint Louis, USA) (CMC). Cellulolytic activity was evidenced by the formation of clear zones through Congo red® (Sigma-Aldrich, CAS: 573-58-0, Saint Louis, USA) staining36.
Molecular identificationFor the molecular characterization, genomic DNA was obtained from each isolate using the DNAzol reagent (INVITROGEN by Thermo Fisher Scientific, Cat. No.10503027, Carlsbad, CA, USA). The ITS rDNA region was amplified using the ITS universal primers ITS1 (5′-TCCGTAGGTGAACCTGCGG-3′) and ITS4 (5′-TCCTCCGCTTATTGATATGC-3′) reported by Cordero-Ramírez et al.9 and White et al.42 PCR was performed in a 25μl volume containing 1ng DNA template, 1.5mM MgCl2, 0.5mM of each dNTP, 0.4μM of forward and reverse primers, and 1U of Taq DNA polymerase (Invitrogen, Brazil, Cat. No. 11615-050). The thermocycler was programmed for initial denaturation at 95°C for 4min, followed by 35 cycles of denaturation at 94°C, annealing at 54°C, extension at 72°C, each for 1min and a final extension at 72°C for 5min PCR products 500bp in length were separated by agarose gel electrophoresis (1% (w/v) in 0.5× TAE) and visualized by ethidium bromide staining. PCR products were purified using the QIAquick PCR purification Kit (Qiagen, Cat. No. 28106) and quantified using a Nanodrop 2000. PCR products were sequenced in both directions with an ABI 3730XL sequencer (Applied Biosystems, USA). Sequences were edited in CHROMAS Pro 1.6 (Technelysium Pty Ltd., South Brisbane, Queensland, Australia) and compared to sequences in the NCBI (National Center for Biotechnology Information) using the BLAST-N software and the Megablast algorithm. The ITS sequences were compared with sequences from the NCBI GenBank database and aligned with 18 reference sequences using the MUSCLE program. Phylogenetic trees were constructed using the Kimura 2-parameter (K2P) model and the Neighbor Joining (NJ) method. The phylogenetic analysis was performed using 1000 bootstraps for astringency in the MEGA 7 software22.
Medium and culture conditionsMineral media A, B, and C were used for hydrolysis and cellulase production, as indicated in the text. The compositions of these media were as follows (g/l): Medium A (MA): 1.0, KH2PO4; 0.7, MgSO4·7H2O; 0.5, NaCl; 0.7, FeSO4; 0.3, NH4NO3; 0.3, MnSO4; pH 4.0. Medium B (MB): 2.0, KH2PO4; 0.4, CaCl2·2H2O; 0.3, MgSO4·7H2O; 0.005, FeSO4·7H2O; 0.0016, MnSO4·H2O; 0.0014, ZnSO4·7H2O; 0.002, CoCl2·6H2O; 0.97, urea; 0.36, yeast extract; pH 5.0, formulation according to Maeda et al.25 Medium C (MC): 2.0, KH2PO4; 0.3, CaCl2.H2O; 0.3 MnSO4·7H2O; 1.4, (NH4)2SO4; 0.005, FeSO4·7H2O; 0.0016, MnSO4·H2O; 0.0014, ZnSO4·7H2O; 0.002, CoCl2·6H2O; 0.25, peptone; 0.75, yeast extract; 0.3, urea; 1.0mL, Tween-80 and pH 5.5, reported by Mandels and Weber27. CMC (1%), Avicel® (Fluka Analytical, Pcode: 101137236, New York, USA) (1%), or moringa milled (0.5mm) straw (2%) were added individually as carbon sources.
All assays were performed in 250ml Erlenmeyer flasks containing 70ml of medium. The flasks were inoculated at a final concentration of 1×106spores/ml and incubated for 168h at 30°C with shaking at 200rpm Samples were collected at different time intervals and centrifuged at 10000rpm for 10min supernatants were stored at −20°C. The experiments were carried out in triplicate.
Reducing sugars and enzyme activity assaysThe concentration of reducing sugars was measured by the 3, 5-dinitrosalicylic acid® (Sigma-Aldrich, CAS: 609-99-4, Saint Louis, USA) (DNS) method29. Cellulolytic activities were determined as previously reported by Zhao et al.44 with some modifications. (EnG) activity was determined by incubating 900μl of CMC (1%) with crude enzymatic extract in a final volume of 1.1ml containing 0.1M acetate buffer, pH 6.0, at 50°C for 50min Filter paper activity (ExG) was evaluated by incubating 12.5mg of filter paper and 800μl of enzyme extract in 1ml of reaction mixture containing 0.6M acetate buffer pH 6.0, at 50°C for 50min For BG activity, 250μl of culture supernatants were added with 250μl of 10mM salicin® (Sigma Life Science, CAS: 138-52-3, Saint Louis, USA) (in 0.1M acetate buffer, pH 6.0) at 50°C for 50min The DNS method was used to measure the sugar content29 for all reactions. One unit of enzymatic activity was defined as the amount of enzyme that liberated 1μmol of reducing sugars (expressed in glucose equivalents) per min.
Statistical analysisStatistical analysis data from the degradation of cellulose substrates and enzymatic assays were subjected to analysis of variance (ANOVA) using the SAS 9.0 program (SAS Institute, Inc., Cary, NC, USA). Duncan's test was used for the post hoc comparison of means (p≤0.05).
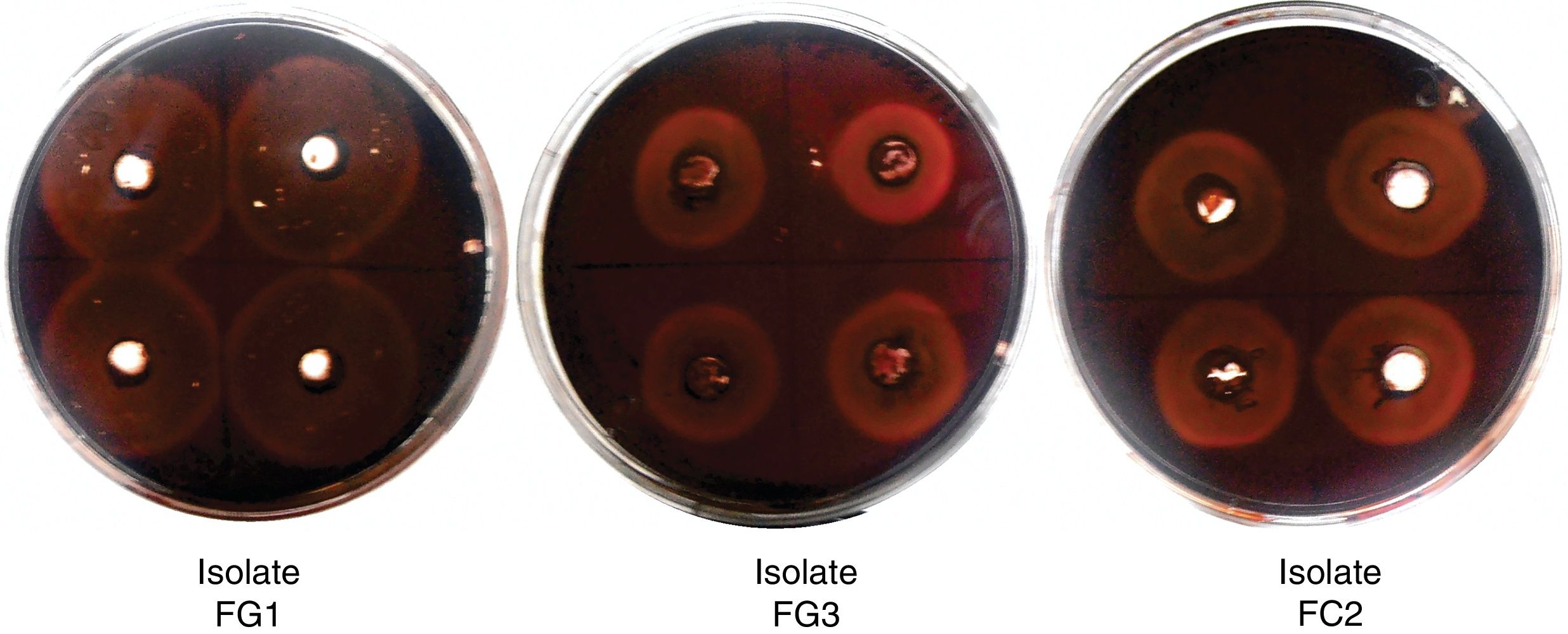
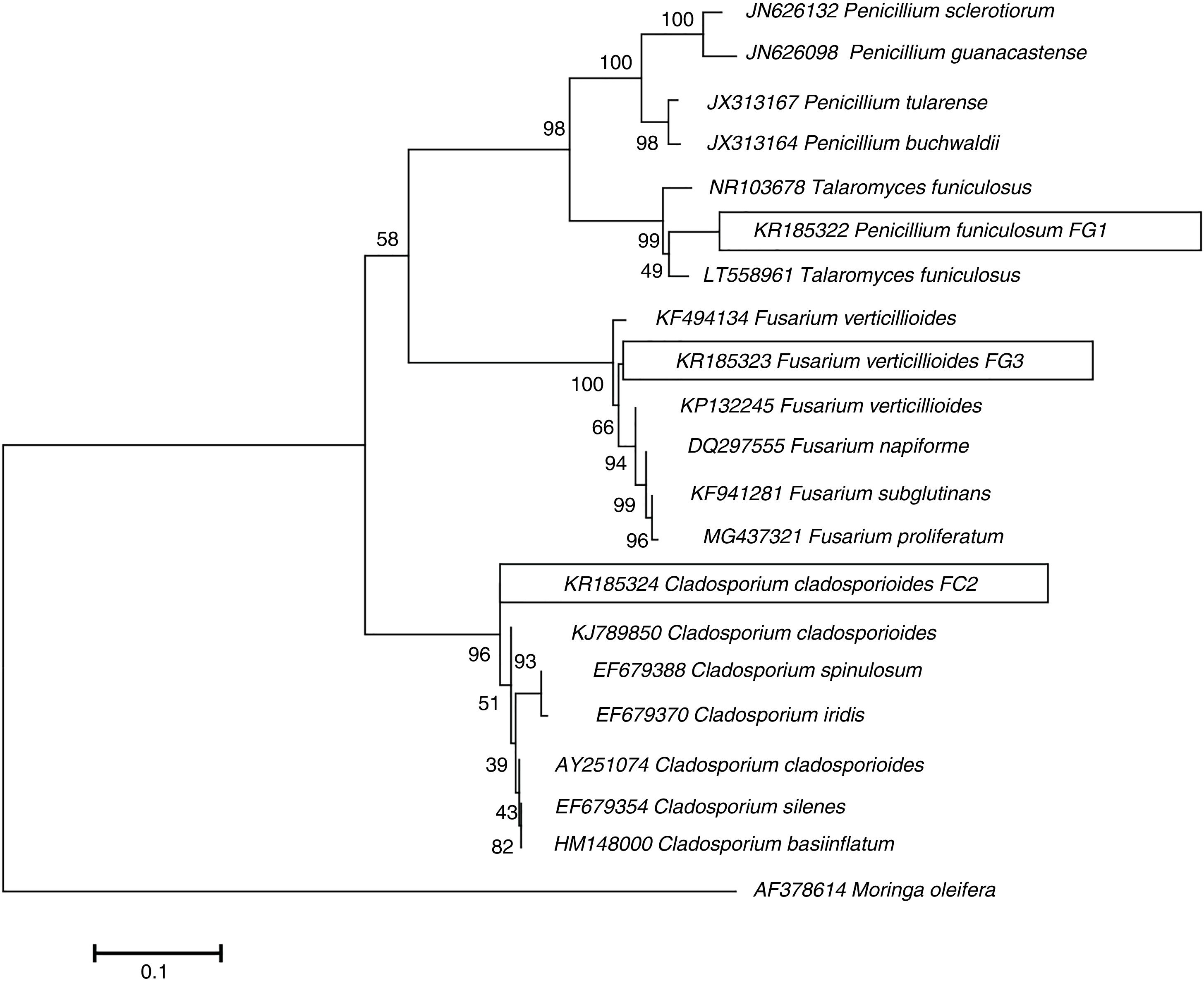
ResultsIsolation and identification of cellulolytic microorganismsIn this study, a total of 120 microorganisms were obtained from M. oleifera biomass; 48 of them were purified (41 bacteria and 7 fungi) and tested for the production of cellulolytic enzymes on CMC-agar plates. The results of the clearing zone method revealed that most of these microorganisms (42) possessed the ability to hydrolyze cellulose. The largest hydrolytic halos were observed for three fungal isolates: FG1, FG3, and FC2 (Fig. 1). For the molecular identification, the DNA of each isolate was analyzed by PCR using the ITS universal primers that amplified fragments of approximately 500bp in size. The ITS sequences were matched with those available in the GenBank datasbase, which revealed the maximum identity of FG1, FG3 and FC2 with the microorganisms Penicillum (Talaromyces funiculosus anamorph: Penicillium funiculosum), Fusariumverticillioides and Cladosporiumcladosporioides respectively (Table 1). The sequences obtained were deposited in the NCBI GenBank under accession numbers KR185322 (Penicillium funiculosum), KR185323 (Fusarium verticillioides), KR185324 (Cladosporium cladosporioides). A phylogenetic tree was constructed using the Neighbor-joining method which showed that each fungal isolate (FG1, FG3 and FC2) was placed in the same clade with their respective species (Fig. 2).
Cellulolytic activity of fungal isolates (qualitative analysis). Agar plate containing CMC as substrate and Congo red as indicator. The wells were filled with the supernatant of the selected strains and incubated at 30°C for 48h. A zone of clearance around the wells indicates cellulose degradation.
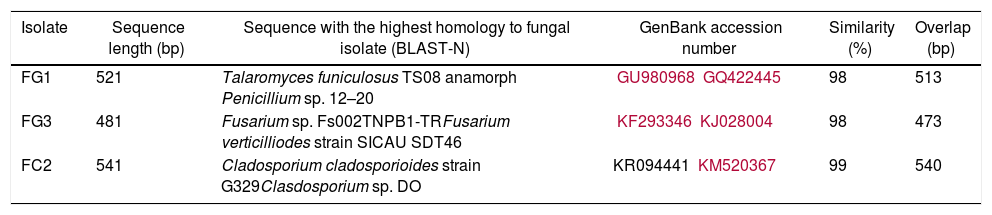
Best matches obtained by BLAST-N (NCBI) using the Megablast algorithm from ITS rDNA sequences obtained from fungal isolates
| Isolate | Sequence length (bp) | Sequence with the highest homology to fungal isolate (BLAST-N) | GenBank accession number | Similarity (%) | Overlap (bp) |
|---|---|---|---|---|---|
| FG1 | 521 | Talaromyces funiculosus TS08 anamorph Penicillium sp. 12–20 | GU980968GQ422445 | 98 | 513 |
| FG3 | 481 | Fusarium sp. Fs002TNPB1-TRFusarium verticilliodes strain SICAU SDT46 | KF293346KJ028004 | 98 | 473 |
| FC2 | 541 | Cladosporium cladosporioides strain G329Clasdosporium sp. DO | KR094441 KM520367 | 99 | 540 |
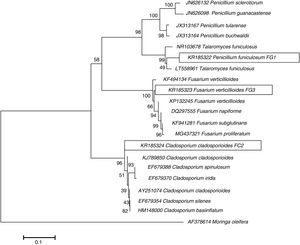
Phylogenetic tree illustrating relationships between the partial sequences (ITS region) of ribosomal genes from isolated fungi and the sequences from strains contained in the GenBank database. The evolutionary history was inferred using the Neighbor-Joining method. The percentage of replicate trees in which the associated taxa clustered together in the bootstrap test (1000 replicates) is shown next to the branches. The tree is drawn to scale, with branch lengths in the same units as those of the evolutionary distances used to infer the phylogenetic tree. The evolutionary distances were computed using the Kimura 2-parameter method and are in the units of the number of base substitutions per site. Evolutionary analyses were conducted in MEGA 7.17.
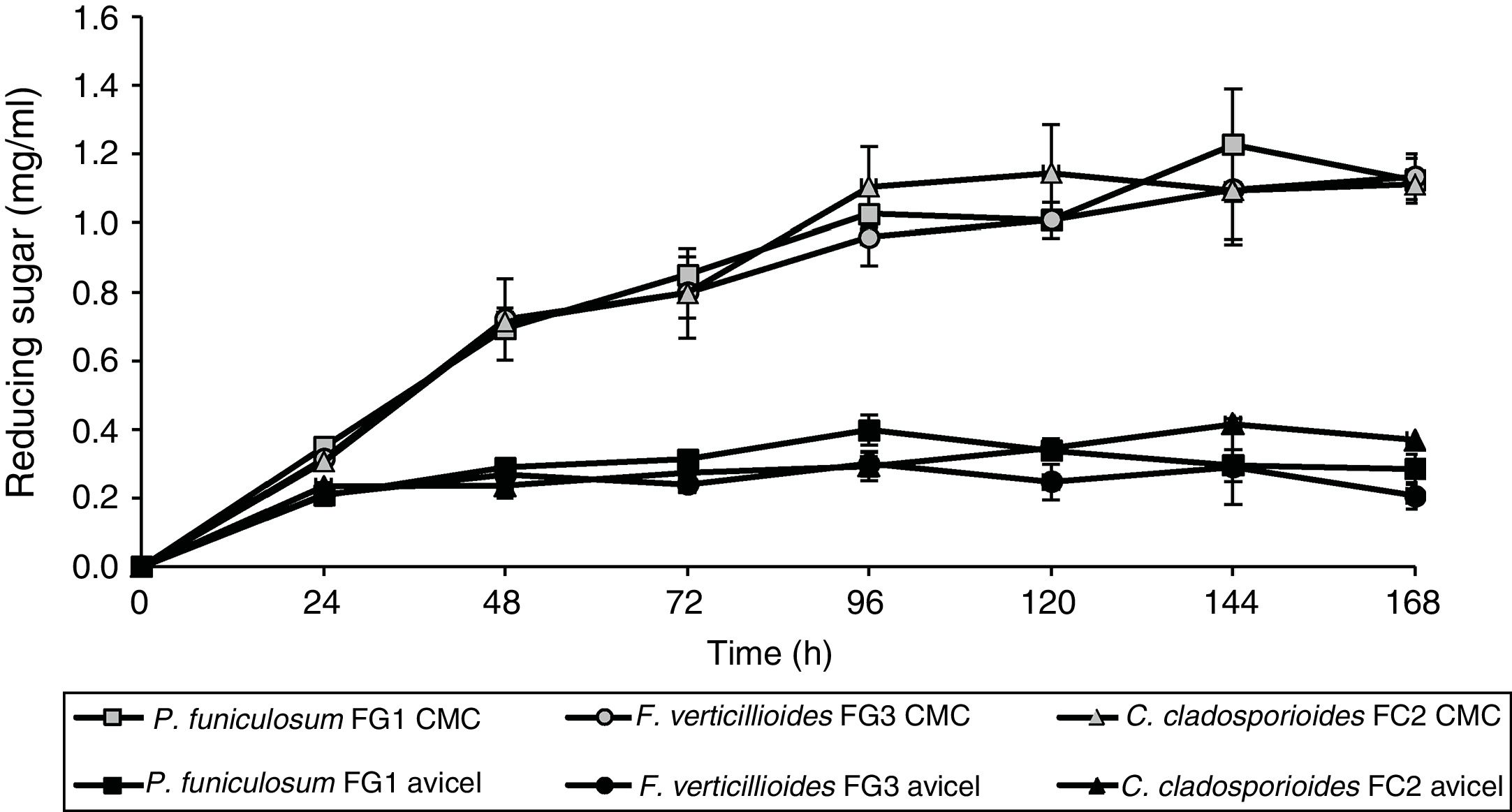
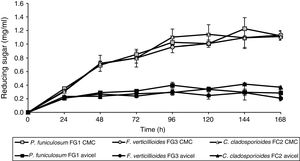
In order to determine their cellulose degradation abilities, P. funiculosum, F. verticillioides, and C. cladosporioides, were grown on medium A, which contained either soluble cellulose (CMC) or the insoluble crystalline form (Avicel) as the sole carbon source. The amounts of reducing sugar released by enzymatic hydrolysis in the different fermentations are shown in Figure 3. The concentration of reducing sugars increased when CMC was the sole carbon source; there was no significant change in the amount of reducing sugars after 24h of fermentation when Avicel was used as the carbon source. The maximum production of reducing sugars was 1.18, 1.14, and 1.13mg/ml for P. funiculosum, C. cladosporioides, and F. verticillioides, respectively, at 96h of culture on CMC. In addition, when the three isolated fungi were cultivated with the same carbon source (either Avicel or CMC), no statistically significant difference was observed between the fungi in terms of reducing sugar production. In contrast, when the strains were cultivated on different carbon sources, significant differences were observed (p < 0.0001).
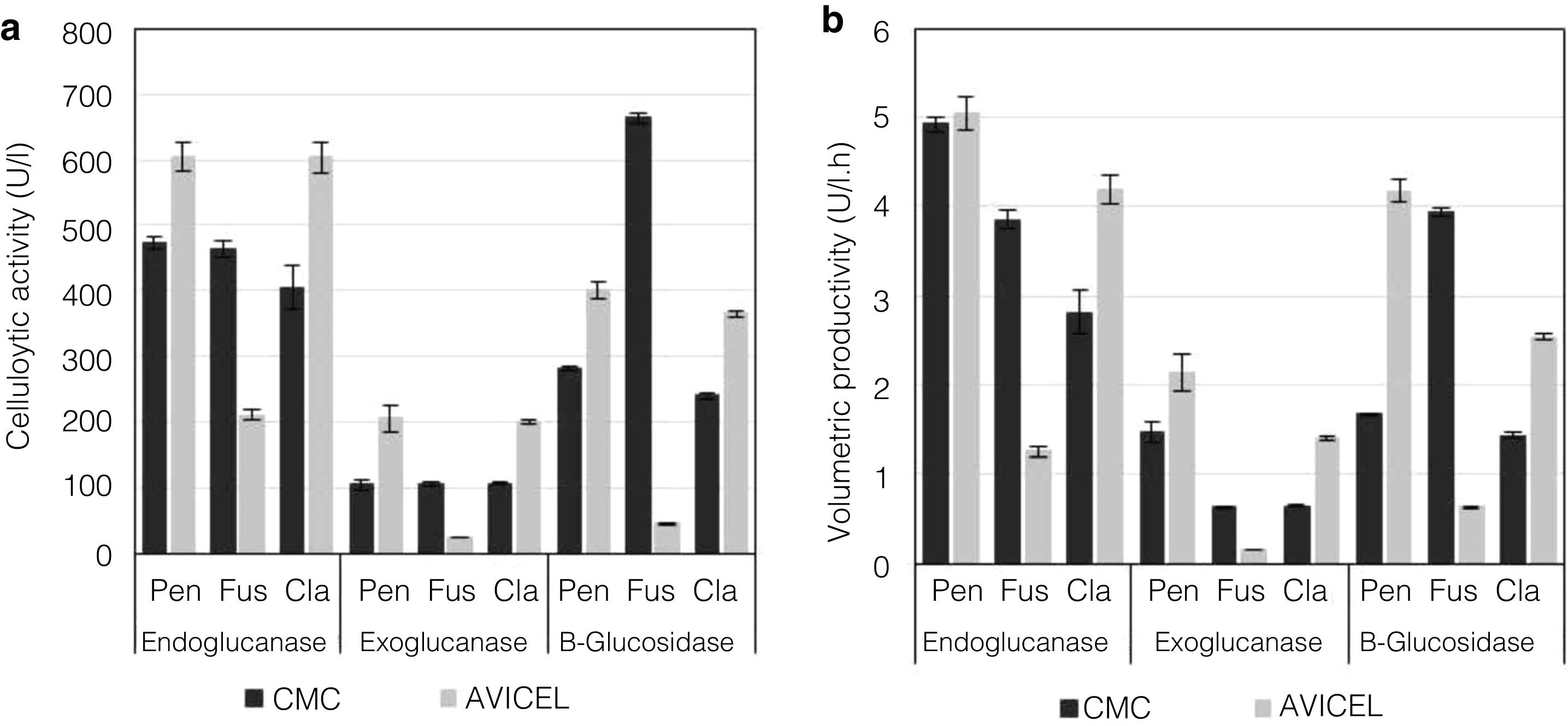
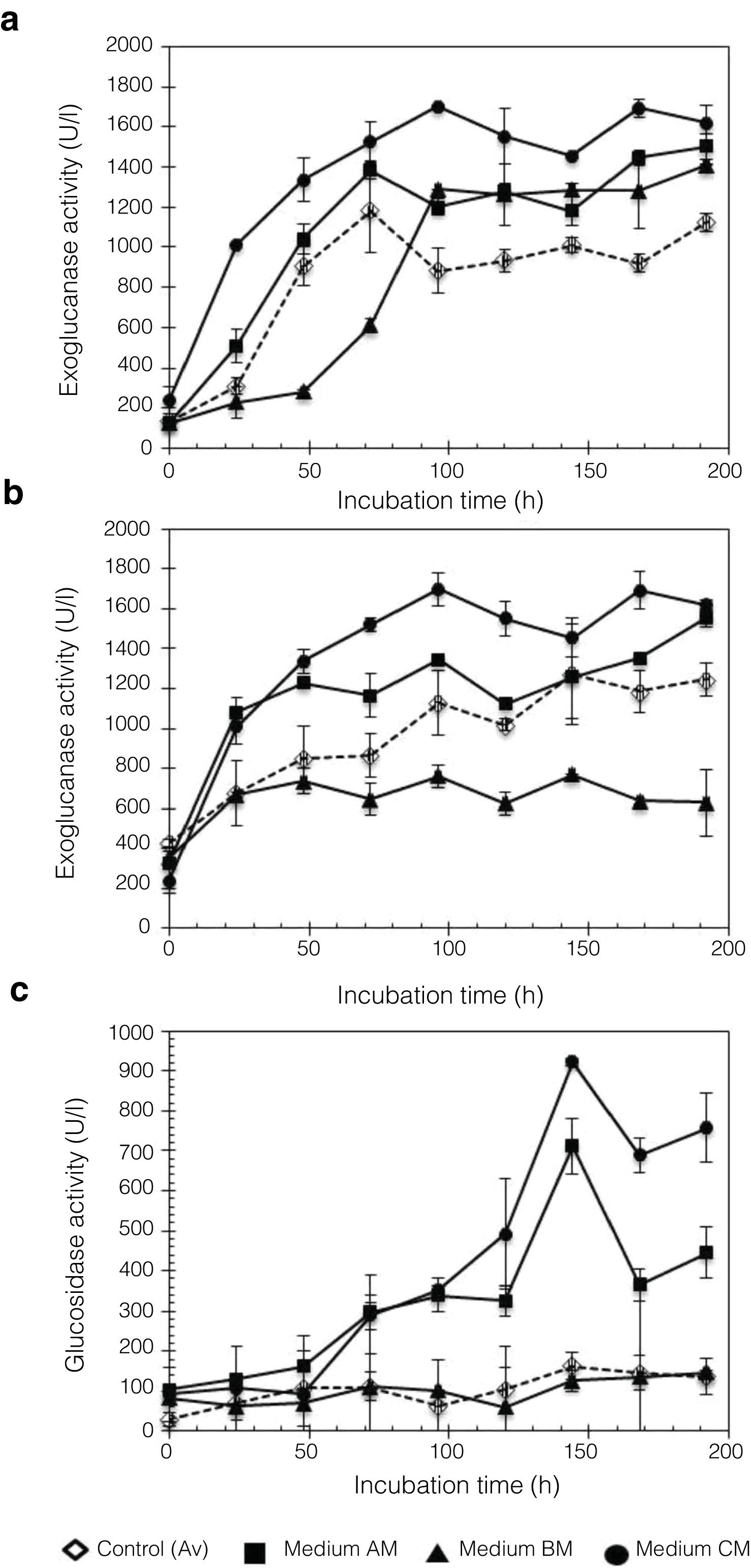
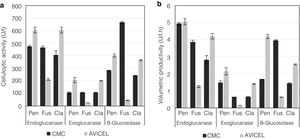
Analysis of enzyme activities in synthetic media as the sole carbon sourceTo evaluate the cellulolytic activities (EnG, ExG, and BG) of the three isolated fungi, the microorganisms were grown on medium “A” containing CMC or Avicel as the sole carbon source. As seen in Figure 4a, the strains presented different enzyme activity on each medium tested. P. funiculosum and C. cladosporioides showed maximum enzyme activity when grown on the Avicel medium, whereas for F. verticillioides the enzymatic activity levels obtained when CMC was used as a sole carbon source were higher than those obtained with Avicel. The highest EnG (606 and 604U/l) and ExG (205 and 200U/l) activities were produced by P. funiculosum and C. cladosporioides, respectively. On the other hand, the maximum BG activity (663.78U/l) was observed for F. verticillioides; although P. funiculosumandC. cladosporioides showed a very similar pattern of cellulolytic enzyme activity. The volumetric productivity revealed that P. funiculosum was more efficient in all cellulolytic activities for either of the two carbon sources (Fig. 4b, Table 2).
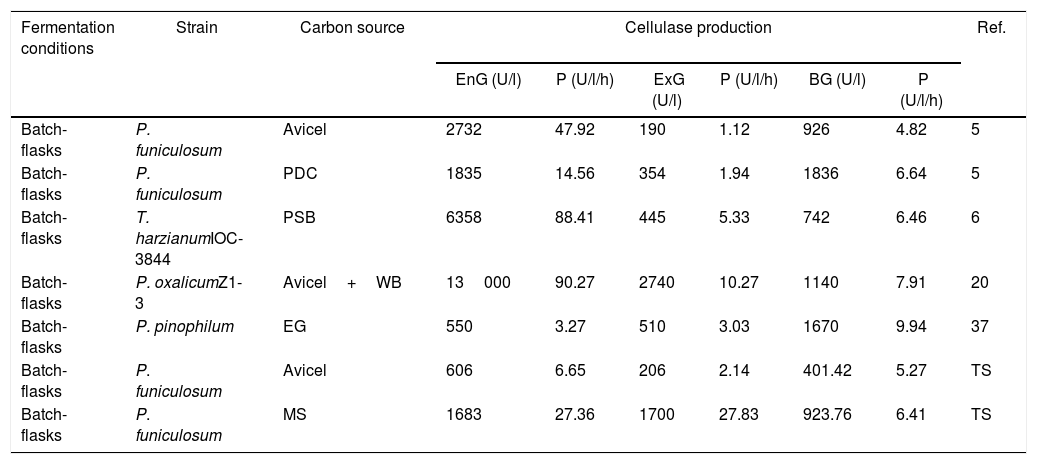
Comparison of cellulolytic enzymatic activities and volumetric productivity of fungi Penicillium, cultivated in submerged fermentation using different carbon sources
| Fermentation conditions | Strain | Carbon source | Cellulase production | Ref. | |||||
|---|---|---|---|---|---|---|---|---|---|
| EnG (U/l) | P (U/l/h) | ExG (U/l) | P (U/l/h) | BG (U/l) | P (U/l/h) | ||||
| Batch-flasks | P. funiculosum | Avicel | 2732 | 47.92 | 190 | 1.12 | 926 | 4.82 | 5 |
| Batch-flasks | P. funiculosum | PDC | 1835 | 14.56 | 354 | 1.94 | 1836 | 6.64 | 5 |
| Batch-flasks | T. harzianumIOC-3844 | PSB | 6358 | 88.41 | 445 | 5.33 | 742 | 6.46 | 6 |
| Batch-flasks | P. oxalicumZ1-3 | Avicel+WB | 13000 | 90.27 | 2740 | 10.27 | 1140 | 7.91 | 20 |
| Batch-flasks | P. pinophilum | EG | 550 | 3.27 | 510 | 3.03 | 1670 | 9.94 | 37 |
| Batch-flasks | P. funiculosum | Avicel | 606 | 6.65 | 206 | 2.14 | 401.42 | 5.27 | TS |
| Batch-flasks | P. funiculosum | MS | 1683 | 27.36 | 1700 | 27.83 | 923.76 | 6.41 | TS |
PDC: sugarcane bagasse partially delignified cellulignin, PSB: pretreated sugarcane bagasse; WB: wheat bran; EG: elephant grass; MS: moringa straw; TS: this study.
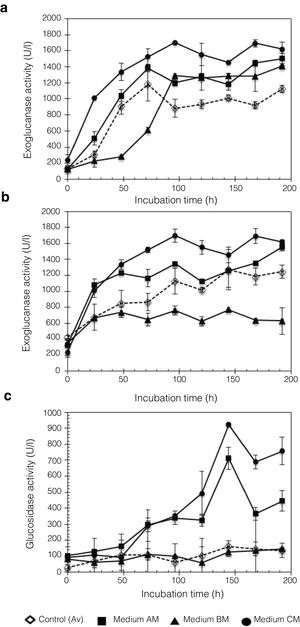
In this work, Penicillium sp. was evaluated using different culture media, A, B, and C, with moringa straw as the sole carbon source. Figure 5 shows the kinetic profile obtained for the cellulolytic enzymes produced by P. funiculosum: EnG, ExG, and BG. In particular, EnG activity reached 1683.27±109.14U/l when medium C containing moringa straw was used, which was 2.77-fold higher than the activity found in A, B, or C without moringa straw (Fig. 5a). Moreover, ExG and BG activities increased by 8.26- and 2.30-fold, respectively, in C medium with moringa straw (Fig. 5b and c). A steady increase in EnG was obtained in the cultivation on medium C from 1313.8U/l to 1624.8U/l during 48–192h of fermentation.
DiscussionIn nature, only a small percentage of microorganisms may degrade cellulose; within these microorganisms, the filamentous fungi are one of the most active and efficient groups due to their capability to secrete different enzymatic activities16. In this study we isolated indigenous cellulolytic microorganisms from decaying biomass of M. oleifera tree. Based on the phylogenetic analysis of the ITS rDNA, the three highest cellulolytic microorganisms were identified as P. funiculosum, F. verticillioides, and C. cladosporoides (Table 1), which have been previously described as cellulase producers by other authors1,4,19,21,41.
When soluble and insoluble cellulose substrates were tested on each isolate, it was evident that reducing sugar levels in the fermentation broth supplied with CMC were higher (approximately 3-fold) than with Avicel. This is possibly due to the fact that CMC is an amorphous material which is easier to be hydrolyzed than Avicel. In addition, differences between the produced hydrolytic enzymes on each studied substrate were noticed. Other authors have reported that the amorphous cellulose regions are preferentially attacked by cellulolytic enzymes, whereas more crystalline areas are resistant to enzymatic attack31,40. Cellulolytic microorganisms use different mechanisms to degrade cellulose in plant cell walls. Most microbes secrete sets of individual cellulases, which act synergistically on native cellulose33,34,39. The analysis of enzyme activities on synthetic media in this study suggests that Avicel is an ideal substrate for the cellulolytic enzymes produced by P. funiculosum and C. cladosporioides, whereas for F. verticillioides the enzymatic activity levels obtained when CMC was used as a sole carbon source were higher than those obtained with Avicel, which is in agreement with the results obtained by Dar et al.10 In addition, P. funiculosum isolated in this study improved productivity of all cellulolytic activities; a similar trend was reported by Carvalho et al.7 who demonstrated that Penicillium funiculosum produced maximum cellulase activities when exposed to Avicel, not CMC or cellobiose. Differences in the cellulose activities produced may be due to multiple factors, such as chemical changes in the culture medium during fermentation (increase and/or decrease of the inducing substance) or by the physiological needs associated with cell growth, as reported by Maeda et al.25 The effect of the culture medium on cellulase production has been studied using different raw materials as the carbon source, including sugarcane bagasse25, wheat bran, corn stover14, and rice straw8. In this work, the moringa straw used as a sole carbon source evidently increased volumetric productivity. Results showed that EnG, ExG, and BG improved by 27.36-, 27.83-, and 6.41-fold, respectively, compared with Avicel (Fig. 5, Table 2). These values suggest that cellulase production was heavily dependent on media composition and cell wall components28. It is clear that Penicillium sp. is a promising strain for production of EnG, ExG, and BG activities. Previous studies have reported that the amount and cooperation between cellulases are crucial in the lignocellulosic biomass hydrolytic process. To date, cellulases used in industry are mainly produced from bacteria and fungi (Penicillium, Aspergillus and Trichoderma). Among them, the filamentous fungus T. reesei is the most important industrial pillar; however, T. reesei has low BG activity, which reduces its efficiency in biomass degradation and compromises its industrial applications12,24,32,33. Our results showed that P. funiculosum was able to produce the three main cellulolytic activities in suitable amounts, showing high BG activity (Fig. 5, Table 2). These results also demonstrated that the cellulolytic activities and volumetric productivity were comparable to other fungal strains (Table 2). However, cellulase productions are still low for the industrial-scale treatment of lignocellulosic biomass. In this sense, it is necessary to continue screening for and characterizing novel cellulase-producing microorganisms using different carbon sources as inducers. Moreover, cellulase activities need to be further enhanced for biofuel applications through the optimization of important process variables.
This study represents the first report on the use of moringa straw as a carbon source for P. funiculosum. This is a cheap nutrient source for the production of cellulolytic enzymes from fungal strains and its use could help to reduce the costs for cellulase production.
Conflict of interestThe authors declare that they have no conflicts of interest.
ELVM acknowledges the Consejo Nacional de Ciencia y Tecnología (CONACYT, México) and the SIP-IPN for Ph.D. fellowships.
This work was financially supported by the Secretaría de Investigación y Posgrado del Instituto Politécnico Nacional (SIP-IPN) (Projects 20131491, 20144369, 20150270, 20160483) and CONACYT, Mexico (Project 2011-16-175519, 270245 and 291143).