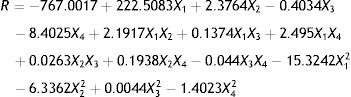Steroids, including testosterone, estrone, 17β-estradiol, estriol and 17β-ethinyl estradiol, are harmful not only to the population dynamics of aquatic life forms but also to public health. In this study, a marine testosterone-degrading bacterium (strain N3) was isolated from Nanao Island in the South China Sea. In addition, the strain could also use 17β-estradiol (E2), 17β-ethinyl estradiol (EE2), estriol (E3) or cholesterol as a sole carbon source. According to the 16S rRNA gene sequence analysis, strain N3 was identified as Vibrio sp. Further characterization showed that the strain is aerobic, gram-negative, and mobile and exhibits resistance to ampicillin, carbenicillin, penicillin and spectinomycin. For enhancing its capacity of testosterone degradation, the Plackett–Burman factorial design and the central composite design were used to optimize the culture condition. Under optimal conditions, 92% of testosterone was degraded by Vibrio sp. N3 in 48h.
Los esteroides—que incluyen la testosterona, la estrona, el 17 β-estradiol, el estriol y el 17 β-etinilestradiol—son nocivos no solo para la población dinámica de las formas de vida acuática, sino también para la salud pública. En este estudio se aisló una bacteria marina degradadora de testosterona de la isla de Nanao, en el Mar del Sur de China, a la que se denominó cepa N3. Se determinó que esta cepa también podría usar 17 β-estradiol (E2), 17 β-etinilestradiol (EE2), estriol (E3) o colesterol como únicas fuentes de carbono. De acuerdo con el análisis de la secuencia del gen 16S rRNA, la cepa N3 se identificó como Vibrio sp. La caracterización adicional mostró que dicha bacteria es un organismo aerobio, gram negativo y móvil, y que presenta resistencia a ampicilina, carbenicilina, penicilina y espectinomicina. Para optimizar la condición de cultivo en relación con su capacidad de degradar la testosterona, se utilizaron el diseño factorial Plackett-Burman y el diseño compuesto central. En condiciones óptimas, el 92% de la testosterona fue degradada por Vibrio sp. N3 en 48h.
Marine pollution has become a global problem in recent years with increasing contamination levels and negative effects in the marine environment. It has been reported that the survival of aquatic organisms and human health are affected by pollution in most coastal areas of the world. The problem of endocrine-disrupting chemicals (EDCs) has emerged as a major environment and human health issue, gaining attention from scientists, governments, and media6. EDCs, including pesticides, plasticizers, and steroids, have been detected in lakes, drinking water, rivers and sea water3,4,13,23,25.
Steroids include natural hormones testosterone, estrone (E1), 17β-estradiol (E2), estriol (E3) as well as artificial hormones such as 17α-ethinylestradiol (EE2). The most likely sources of testosterone are municipal sewage sludge, paper mill effluents, and runoff from agricultural production sites14,15,19. Steroid hormones are able to act as endocrine disruptors in aquatic environments, and are harmful to humans and other mammals. For example, it has been shown that wild fish living downstream of municipal sewage draining exits were negatively affected by androgenic steroids26.
Several environmental models such as soils8,20, waste water treatment plants10, and stream sediments have been used to degrade testosterone7. Furthermore, the application of microorganisms to degrade testosterone is a promising technological choice. It has been reported that terrestrial bacteria can degrade testosterone completely into carbon dioxide and water11,12,16. In addition, two marine bacteria isolated from the Baltic Sea at Kiel in Germany were found to be able to degrade testosterone, estradiol, and cholesterol22,27.
Statistics-based experimental designs have proved useful in the optimization of biotreatment processes2,28. The Plackett–Burman design (PBD) is widely used to screen variables that have a significant impact on the process13. Response surface methodology is an empirical modeling technique to evaluate the relationship between a set of controlled experimental factors and observed results1.
In this study, four testosterone-degrading strains N1, N3, N6 and N13 were isolated from sediment in the marine fish culture area near Nanao Island, South China Sea. Its taxonomic characterization and steroid degradation capacity were analyzed. The optimal testosterone degradation condition for strain N3 was established.
Materials and methodsChemicals and culture mediaChemicals (analytical grade), including cholesterol, testosterone, 17β-estradiol, estrone, estriol, and 17α-ethinyl-estradiol, were purchased from Sigma (USA).
The mineral medium (MM) was prepared with the following ingredients: 1.0g/l (NH4)2SO4, 0.8g/l Na2HPO4, 0.2g/l KH2PO4, 0.2g/l MgSO4, 1ml/l FeCl3(sq), 1m/l (NH4)6Mo7O24(sq), and 25g/l NaCl. The 2216E medium was prepared with 5g/l peptone, 1g/l yeast extract, 0.1g/l FePO4, and 1l aged seawater.
Testosterone quantification with HPLCTestosterone was extracted with the same volume of ethyl acetate, dried with nitrogen gas, and dissolved in 1ml HPLC elution buffer (ethyl acetate:H2O in 1:1); 10μl testosterone solution was injected into a Hypersil ODS2 C18 4μm×250μm column. The detection program was carried out at flow speed 1ml/min at 30°C for 15min. The testosterone peak was observed after 6.745min at 245nm.
Isolation of steroid-degrading strainsThe strains were isolated from sediment in the marine fish culture area near Nanao Island, South China Sea. Newly collected seabed sludge (5g) and surface seawater (50ml) were thoroughly mixed in a shake flask, cultured in a rotary shaker (120rpm) at 25°C for 30min, and allowed to settle for 30min. The cleaner upper phase was centrifuged at 5000×g for 20min, the supernatant was discarded, and the pellet was resuspended in 0.5ml surface seawater. A serial 10-fold dilution was prepared with physiological saline, and 0.1ml dilutions were spread on MM agar plates, containing 10mg/l testosterone as the carbon source. Plates were incubated at 25°C for 3 days. The colonies were streaked repeatedly until morphologically distinct colonies were generated. The colonies were inoculated in MM medium containing 10mg/l testosterone as the sole carbon source and shake-incubated. The cultures with increased turbidity (absorbance at 600nm) were selected for further analysis.
Testosterone degradation by selected bacteriaThe capacity of selected strains to degrade testosterone was further studied in two means. The first was carried out as follows: the selected strains (OD600 of 0.01) were cultured with 0.3mM testosterone in a 1:10 diluted low carbon source 2216E medium for 3 days. The culture without testosterone was used as a control to determine bacterial growth. Total protein was determined with the Bradford method (subtracting the protein of control). After cultivation, an equal volume of acetonitrile was mixed with the culture solutions for 30min at room temperature. After centrifugation at 12000×g for 20min, the supernatant was transferred into new tubes, and testosterone concentrations were measured by HPLC.
For the second approach, selected bacteria were cultured in 2216E medium at 25°C overnight, collected by centrifugation (12000×g, 5min), and washed twice with MM medium. Pellets were resuspended in MM medium to an OD600 of 0.5, and testosterone (0.3mM) was added as the sole carbon source. Cultures were incubated in a rotary shaker (150rpm) at 25°C for 24h. The OD600 of the cultures was measured every 6h. The remaining testosterone in the cultures was quantified by HPLC.
Identification of bacterial strainThe overnight culture of N3 cells (500μl) was mixed with 50μl SDS (10%) at room temperature for 2min. Then, 300μl phenol and 300μl chloroform were used to extract the chromosomal DNA of the cells twice. After precipitation with 1ml ethanol and 25μl NaCl (5M), the DNA was centrifuged and washed with 70% ethanol twice. Finally, the DNA was resuspended in 50μl TE buffer. The purified DNA was used as template for PCR.
The 16S rRNA gene of strain N3 was amplified by PCR with primers 5′-AGAGTTTGATCCTGGCTCAG-3′ and 5′-GGTTACCTTGTTACGACTT-3′. The PCR product (1489bp) was introduced into pMD-19T (Takara) and the plasmid was transformed into E. coli DH-5α. Three positive clones were further verified by sequencing.
The obtained 16S rRNA gene sequence was subjected to similarity search on the NCBI web site. The highly similar 16S rRNA gene sequences of related species were obtained. Multiple sequence alignment was carried out by using the Clustal X program (version 1.8) with default parameters. Phylogenetic and distance analysis of the aligned sequences were performed by using the MEGA software (version 2.1). The resulting unrooted tree topologies were evaluated by bootstrap analysis of the neighbor-joining method based on 1000 resampling.
The identification of the N3 strain was ascertained by physiological and biochemical tests9.
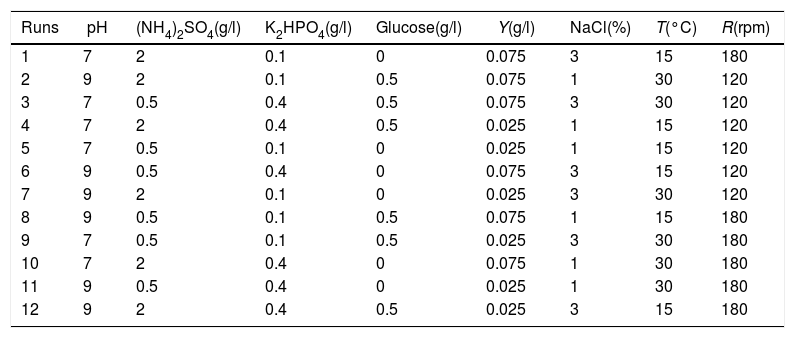
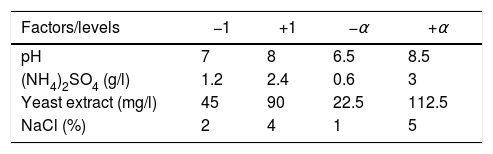
Plackett–Burman design for main factor screeningPlackett–Burman designs were used for screening the fertilization factors that affected testosterone degradation rate. Eight factors (pH, culture temperature (T), shaker roll-rate (R), (NH4)2SO4, K2HPO4, glucose, NaCl, and yeast extract (Y)) were chosen as variables in the study. The degradation rates of testosterone were evaluated after 72h of the culture in each condition containing the same amount of testosterone. The design and analysis of the experiments were performed using the Minitab 7.0 program. Each experiment was carried out three times. The levels of each factor are given in Table 1.
Level setting and designation of the Plackett–Burman experimental design
| Runs | pH | (NH4)2SO4(g/l) | K2HPO4(g/l) | Glucose(g/l) | Y(g/l) | NaCl(%) | T(°C) | R(rpm) |
|---|---|---|---|---|---|---|---|---|
| 1 | 7 | 2 | 0.1 | 0 | 0.075 | 3 | 15 | 180 |
| 2 | 9 | 2 | 0.1 | 0.5 | 0.075 | 1 | 30 | 120 |
| 3 | 7 | 0.5 | 0.4 | 0.5 | 0.075 | 3 | 30 | 120 |
| 4 | 7 | 2 | 0.4 | 0.5 | 0.025 | 1 | 15 | 120 |
| 5 | 7 | 0.5 | 0.1 | 0 | 0.025 | 1 | 15 | 120 |
| 6 | 9 | 0.5 | 0.4 | 0 | 0.075 | 3 | 15 | 120 |
| 7 | 9 | 2 | 0.1 | 0 | 0.025 | 3 | 30 | 120 |
| 8 | 9 | 0.5 | 0.1 | 0.5 | 0.075 | 1 | 15 | 180 |
| 9 | 7 | 0.5 | 0.1 | 0.5 | 0.025 | 3 | 30 | 180 |
| 10 | 7 | 2 | 0.4 | 0 | 0.075 | 1 | 30 | 180 |
| 11 | 9 | 0.5 | 0.4 | 0 | 0.025 | 1 | 30 | 180 |
| 12 | 9 | 2 | 0.4 | 0.5 | 0.025 | 3 | 15 | 180 |
R: shaker roll-rate; T: culture temperature; Y: yeast extract.
Response surface methodology with a central composite design was used for the main factors of these parameters on testosterone degradation using the Design Expert 7.0 program. Based on the results of the Plackett–Burman experiment, four factors (pH, (NH4)2SO4, NaCl, yeast extract) were chosen as the main factors for further condition optimization. Therefore, a four-factor response surface methodology with a central composite design was constructed. The design had eight star points and six replicates at center points leading to 30 runs. The independent factors along with their levels are shown in Table 2. Testosterone degradation rates were measured after a 72h culture period.
Results and discussionIsolation of testosterone-degrading bacteriaA total of 13 bacteria strains were isolated in this study and marked as N-1 to N-13 respectively. The preliminary study had shown that strains N1, N3, N6 and N13 could grow up in MM medium with 10mg/l testosterone as the sole carbon source, which indicates they have the potential to degrade; therefore, strains N1, N3, N6 and N13 were selected for further research.
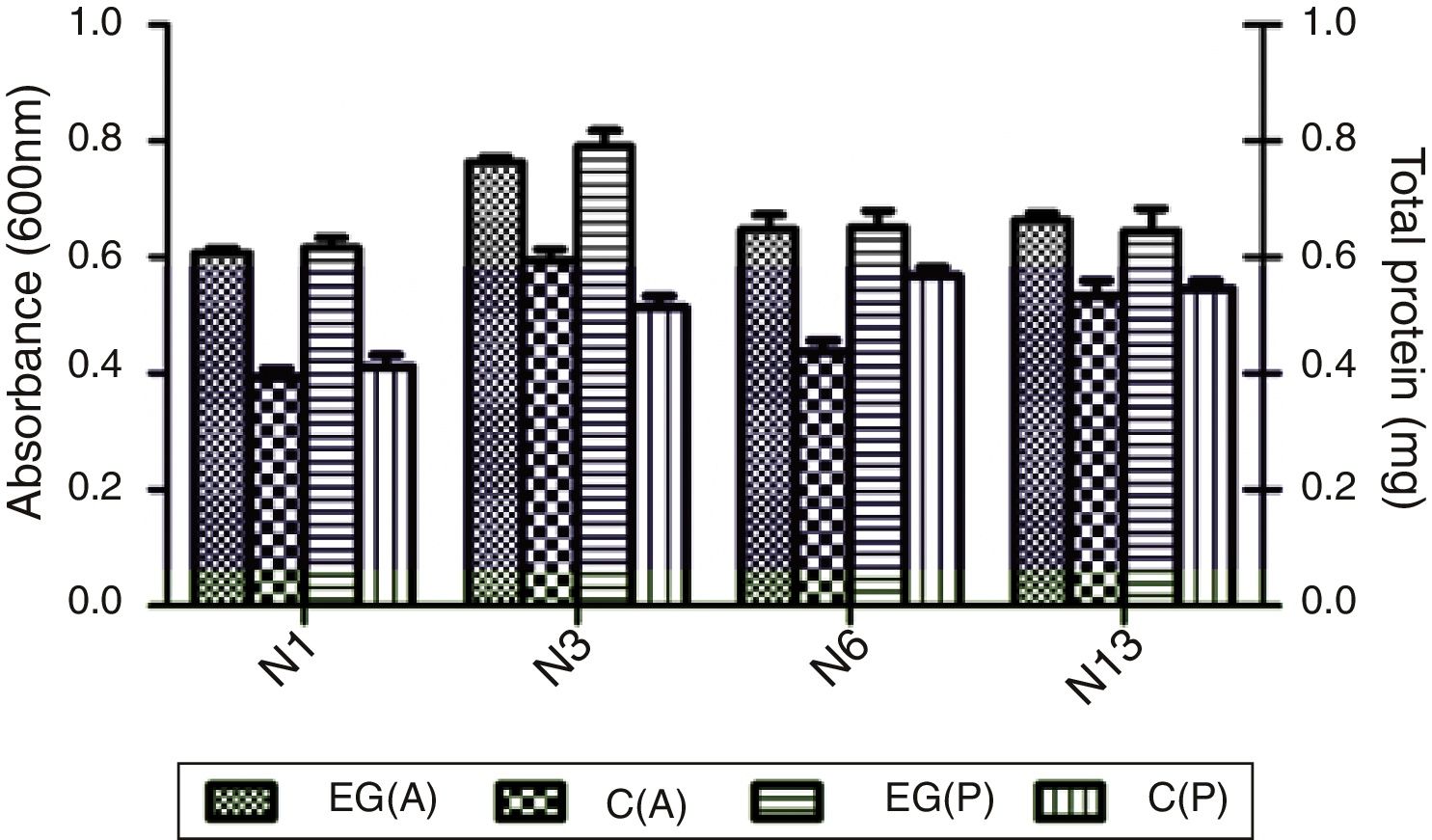
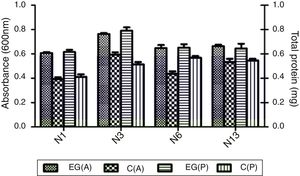
Degradation of testosterone by the selected strainsCompared to the control medium which contained no carbon source, all four strains, N1, N3, N6 and N13, grew as shown in OD600, and produced more total proteins in the medium supplemented with 0.3mM testosterone (Fig. 1). The result indicated that strains N1, N3, N6 and N13 could use testosterone as carbon source. The result was in accordance with the marine, steroid-degrading bacterium H5, which showed better growth in cultures including testosterone, estradiol and cholesterol compared to the non-steroid controls27. Among the four strains, N3 could reach the highest concentration and produce a high amount of total proteins.
Absorbance (A) at 600nm and total protein (P) of four selected strains N1, N3, N6, and N13 cultivated in 1:10-diluted 2216E media for 3 days. EG(A) and EG(P) are the absorbance and protein content of the experiment group in which testosterone was added to media with a concentration of 0.3mM. C(A) and C(P) are the absorbance and protein content of the control group without any additional carbon.
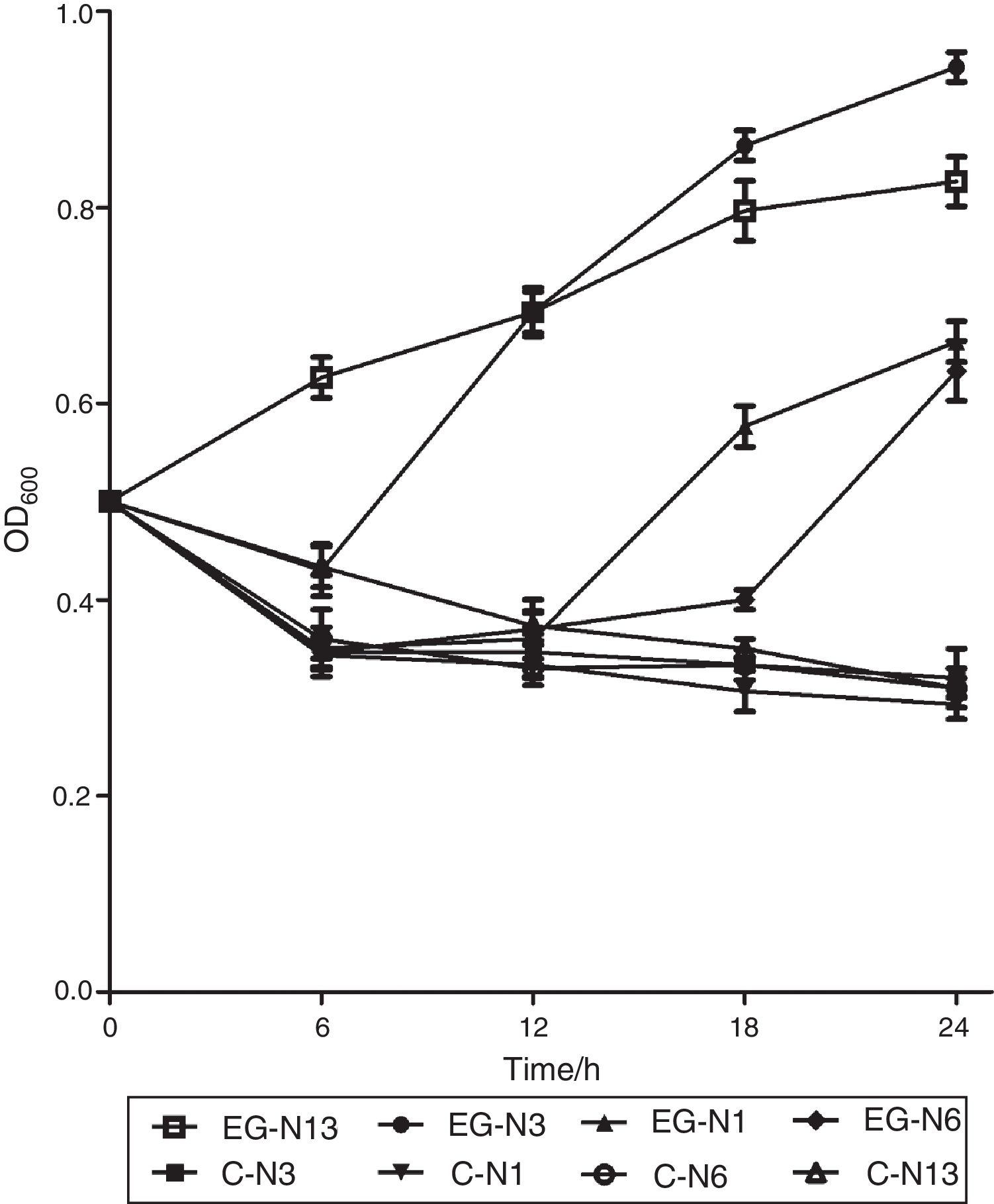
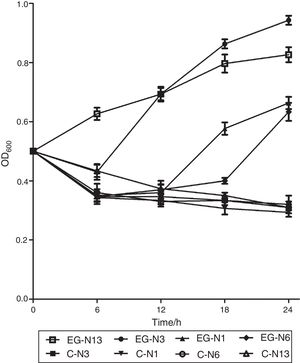
Absorbance at 600nm of strain N13 increased gradually to about 0.85 during the 24h period (Fig. 2). However, absorbance decreased initially at 6, 12 and 18h for strain N3, N1 and N6, and then increased to about 1.0, 0.68, 0.65 at 24h, respectively (Fig. 2). For strain N1, N3, N6, N13, during the three day-incubation, the degradation rate of testosterone was 46.39%, 53.61%, 37.48% and 50.79%, respectively. Strain N3 showed the highest testosterone degradation rate. Moreover, strain N3 could also degrade a wide range of steroids such as E1, E2, E3, EE2, and cholesterol (data no shown), although not as efficiently as the degrading testosterone. Estrone and testosterone have been reported to be the two main steroid pollutants in the environment5. Therefore, strain N3 was chosen for further research.
Identification and characterization of strain N3The results of the characterization showed that strain N3 is aerobic, gram negative and mobile. The result from the Methyl Red-VP test (MR-VP) was negative (data not shown); suggesting that strain N3 does not express pyruvate decarboxylase. N3 could not use citrate as the sole carbon source but could degrade proline into carbon dioxide and water. The strain is resistant to ampicillin (200μg/ml), carbenicillin (200μg/ml), penicillin (150μg/ml) and spectinomycin (30μg/ml); but sensitive to chloromycetin (6μg/ml), tetracycline (6μg/ml), and rifampicin (6μg/ml). Strain N3 can survive under low concentrations (6μg/ml) of kanamycin, streptomycin, and nalidixic acid.
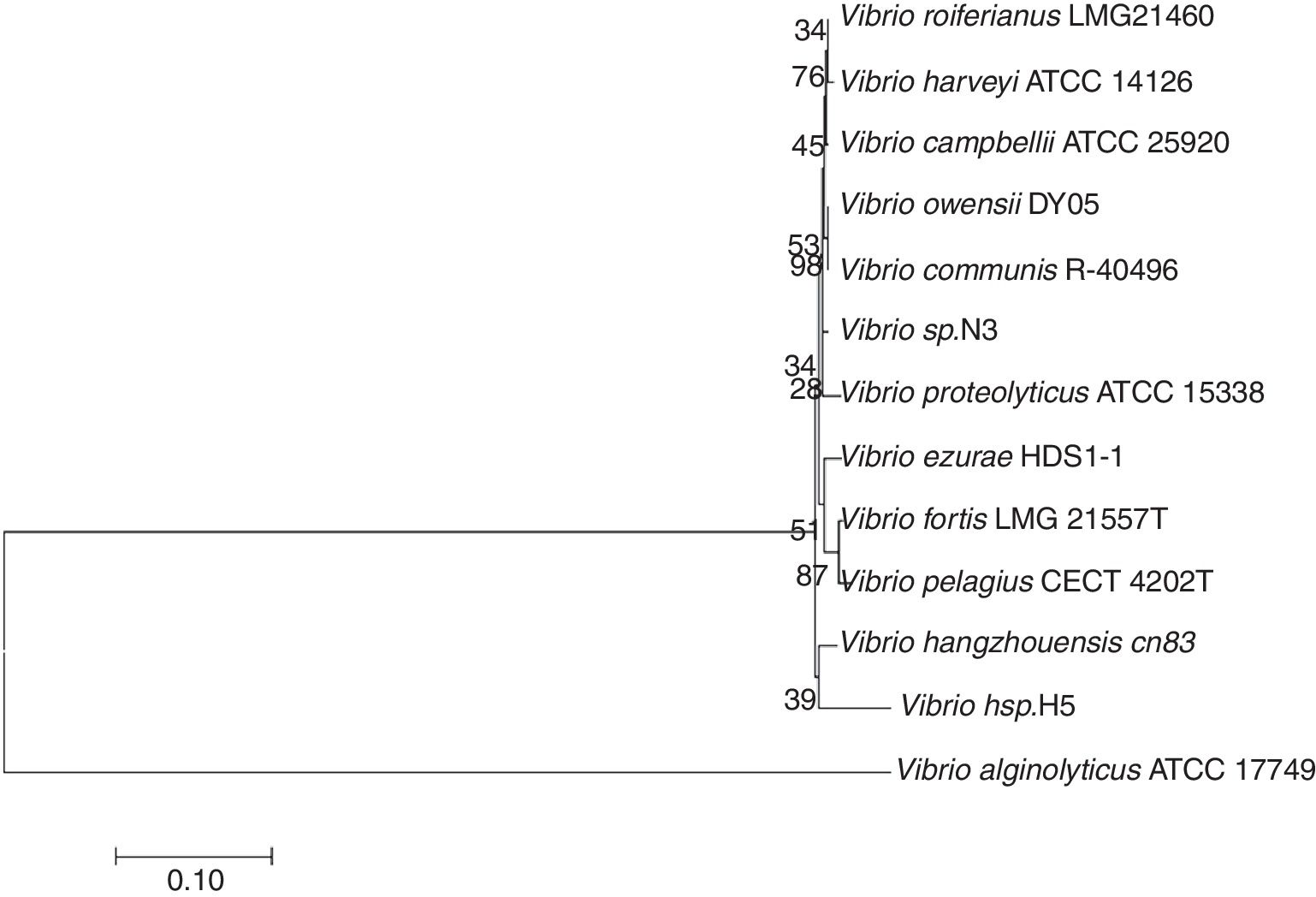
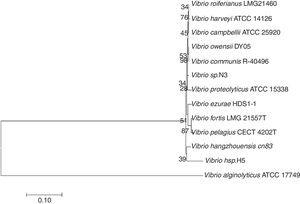
The 16S rRNA gene was amplified from the total DNA of strain N3 and sequenced. The accession number of the 16S rRNA gene of strain N3 in GenBank is JN402325. After alignment with other 16S rRNA gene sequences in NCBI, the result showed high similarity (>97%) to members of the genus Vibrio. The 16S rRNA gene sequence was further aligned with the corresponding sequences from additional strains of Vibrio species in Bergey's Manual of Determinative Bacteriology, which were retrieved from GenBank (Fig. 3). Vibrio sp. H5 is the first bacterium isolated from the marine environment, which is able to degrade testosterone22,27. The sequence similarity of the 16S rRNA gene between strain H5 and strain N3 is 91%, which indicates that Vibrio sp. N3 may represent a new species of Vibrio that could use steroids as the sole carbon source (Fig. 3).
Screening the factors affecting testosterone degradationCulture temperature (T), pH and shaker roll-rate (R) may affect the physiological and biochemical conditions of the organism. (NH4)2SO4 and K2HPO4 are necessary nutrients, and can supply the organism with macro elements nitrogen, phosphorus, and sulfur. It has been reported that low concentrations of glucose could enhance phenol degradation because glucose intake could facilitate bacteria uptake of the substrate21. The bacterium was isolated from the sea, which was highly saline. Different concentrations of NaCl were added to the culture medium, because NaCl was selected as an effect factor. Yeast extract contains certain amino acids, peptides, vitamins, and several organic acids that are important to the organism23–25. Eight factors (pH, culture temperature (T), shaker roll-rate (R), (NH4)2SO4, K2HPO4, glucose, NaCl, and yeast extract (Y)) were chosen as variables in the study. For exploring the effect of pH, culture temperature (T), shaker roll-rate (R), (NH4)2SO4, K2HPO4, NaCl, glucose, and yeast extract (Y) to testosterone degradation, a Plackett–Burman design was used to identify the main effect factors. Results of variable analysis are shown in Table 3.
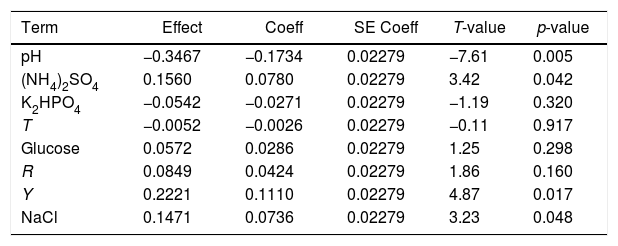
Analysis of variables in the regression model of the Plackett–Burman design
| Term | Effect | Coeff | SE Coeff | T-value | p-value |
|---|---|---|---|---|---|
| pH | −0.3467 | −0.1734 | 0.02279 | −7.61 | 0.005 |
| (NH4)2SO4 | 0.1560 | 0.0780 | 0.02279 | 3.42 | 0.042 |
| K2HPO4 | −0.0542 | −0.0271 | 0.02279 | −1.19 | 0.320 |
| T | −0.0052 | −0.0026 | 0.02279 | −0.11 | 0.917 |
| Glucose | 0.0572 | 0.0286 | 0.02279 | 1.25 | 0.298 |
| R | 0.0849 | 0.0424 | 0.02279 | 1.86 | 0.160 |
| Y | 0.2221 | 0.1110 | 0.02279 | 4.87 | 0.017 |
| NaCl | 0.1471 | 0.0736 | 0.02279 | 3.23 | 0.048 |
coeff: coefficient; p-value: hypothesis tests; R: shaker roll-rate; SE coeff: standard error coefficient; T: culture temperature; t-value: Student's t-test; Y: yeast extract.
Four of the eight factors, including initial pH, yeast extract, (NH4)2SO4, and NaCl, had a significant effect on testosterone degradation (p<0.05), and of these four factors, initial pH showed the most significant effect (p<0.01). Therefore, these four factors were chosen for further optimization. Temperature (T), shaker roll-rate (R), K2HPO4, and glucose were less effective on testosterone degradation by strain N3, and based on the results of the experiment, the highest degradation rate of testosterone was obtained under the condition of 25̊C, 0.025g/l glucose, a shaker roll rate of 150rpm, and 0.25g/l K2HPO4.
Response surface methodology for optimizing the conditions of testosterone degradation of strain N3A central composite design was applied to analyze the main and interactive effects of the four significant factors and to find the optimum values within a coded range of −α to +α in the testosterone degradation rate.
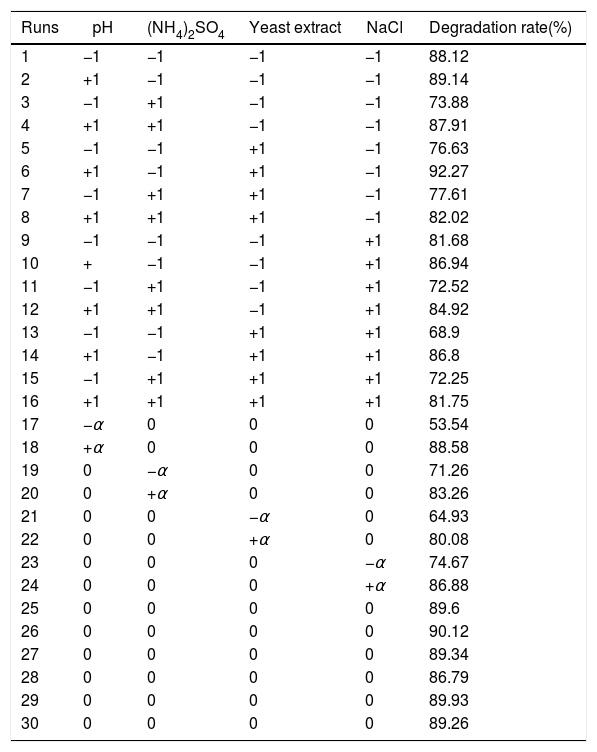
The experimental design and results are shown in Table 4. The experiment results were modeled with an equation to explain the relationship between the testosterone degradation rate and each of these four factors. The mathematical regression model using the coded factors is given as:
where R is the predicted response of the testosterone degradation rate and X1 is the initial pH, and X2, X3, and X4 are concentrations of (NH4)2SO4, yeast extract, and NaCl, respectively.Biodegradation rate of testosterone by Vibrio sp. N3 expose to different factors
| Runs | pH | (NH4)2SO4 | Yeast extract | NaCl | Degradation rate(%) |
|---|---|---|---|---|---|
| 1 | −1 | −1 | −1 | −1 | 88.12 |
| 2 | +1 | −1 | −1 | −1 | 89.14 |
| 3 | −1 | +1 | −1 | −1 | 73.88 |
| 4 | +1 | +1 | −1 | −1 | 87.91 |
| 5 | −1 | −1 | +1 | −1 | 76.63 |
| 6 | +1 | −1 | +1 | −1 | 92.27 |
| 7 | −1 | +1 | +1 | −1 | 77.61 |
| 8 | +1 | +1 | +1 | −1 | 82.02 |
| 9 | −1 | −1 | −1 | +1 | 81.68 |
| 10 | + | −1 | −1 | +1 | 86.94 |
| 11 | −1 | +1 | −1 | +1 | 72.52 |
| 12 | +1 | +1 | −1 | +1 | 84.92 |
| 13 | −1 | −1 | +1 | +1 | 68.9 |
| 14 | +1 | −1 | +1 | +1 | 86.8 |
| 15 | −1 | +1 | +1 | +1 | 72.25 |
| 16 | +1 | +1 | +1 | +1 | 81.75 |
| 17 | −α | 0 | 0 | 0 | 53.54 |
| 18 | +α | 0 | 0 | 0 | 88.58 |
| 19 | 0 | −α | 0 | 0 | 71.26 |
| 20 | 0 | +α | 0 | 0 | 83.26 |
| 21 | 0 | 0 | −α | 0 | 64.93 |
| 22 | 0 | 0 | +α | 0 | 80.08 |
| 23 | 0 | 0 | 0 | −α | 74.67 |
| 24 | 0 | 0 | 0 | +α | 86.88 |
| 25 | 0 | 0 | 0 | 0 | 89.6 |
| 26 | 0 | 0 | 0 | 0 | 90.12 |
| 27 | 0 | 0 | 0 | 0 | 89.34 |
| 28 | 0 | 0 | 0 | 0 | 86.79 |
| 29 | 0 | 0 | 0 | 0 | 89.93 |
| 30 | 0 | 0 | 0 | 0 | 89.26 |
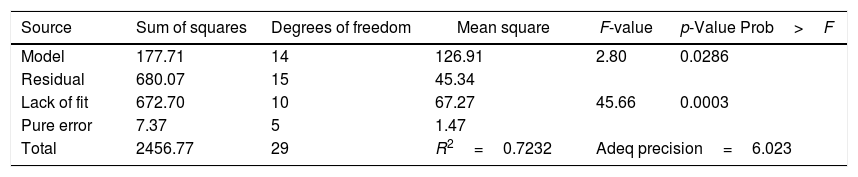
The experiment results of the regression model analysis are shown in Table 5. The regression coefficients and the interaction between each independent factor were considered statistically significant for p-value of 0.0286 (p<0.05). R2 correlation coefficient of the quadratic regression model was 0.7232. The Model F-value of 2.80 implied the model was significant. There was only a 2.86% chance that such a large “Model F-Value” could occur due to noise.
Regression analysis for the biodegradation of testosterone by Design Expert 7.0 for quadratic response surface model fitting (ANOVA)
| Source | Sum of squares | Degrees of freedom | Mean square | F-value | p-Value Prob>F |
|---|---|---|---|---|---|
| Model | 177.71 | 14 | 126.91 | 2.80 | 0.0286 |
| Residual | 680.07 | 15 | 45.34 | ||
| Lack of fit | 672.70 | 10 | 67.27 | 45.66 | 0.0003 |
| Pure error | 7.37 | 5 | 1.47 | ||
| Total | 2456.77 | 29 | R2=0.7232 | Adeq precision=6.023 | |
Adeq precision measures the signal-to-noise ratio, and a ratio greater than 4 is desirable. The ratio of the model was 6.023, indicating that this model could be used to navigate the design.
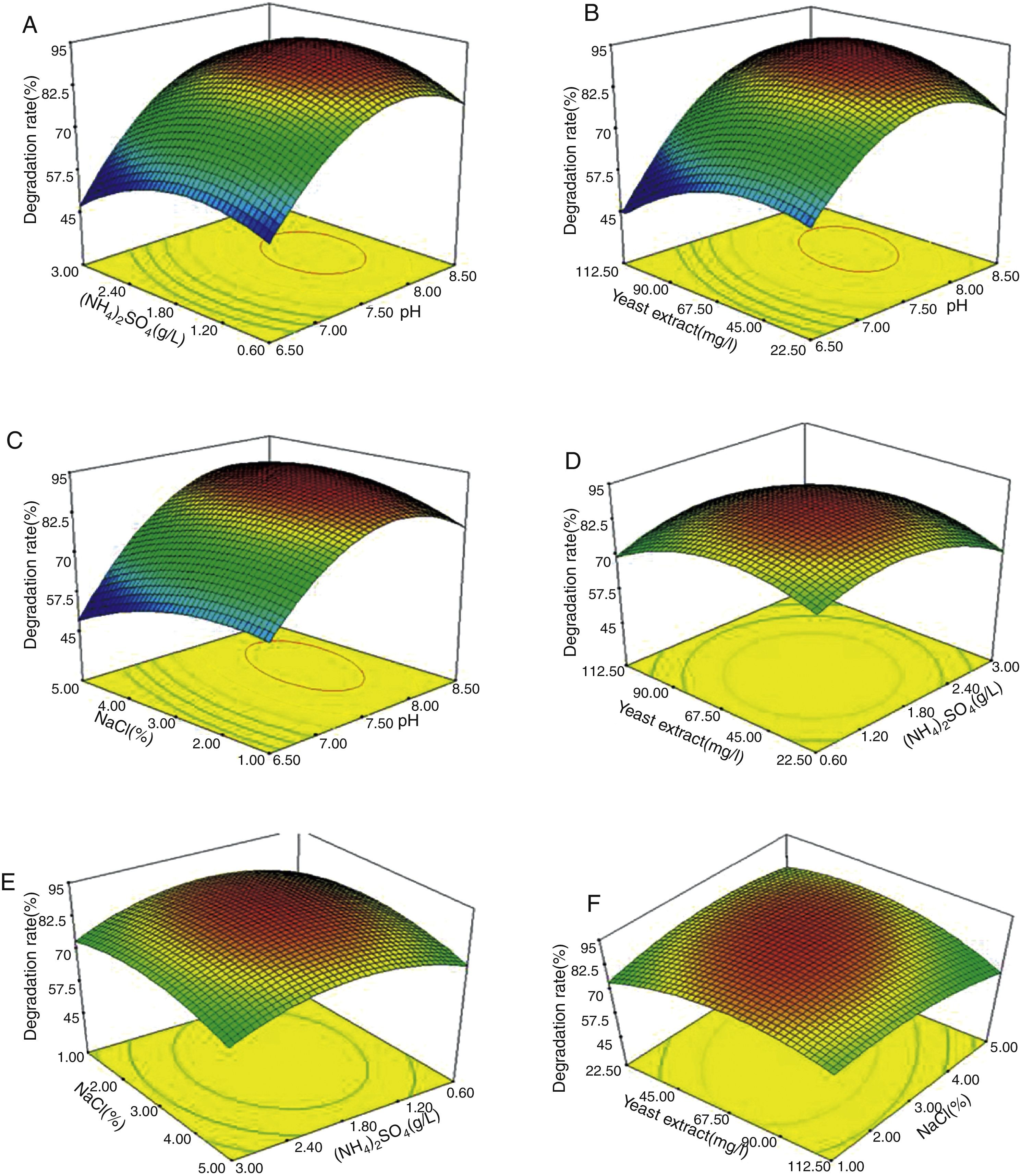
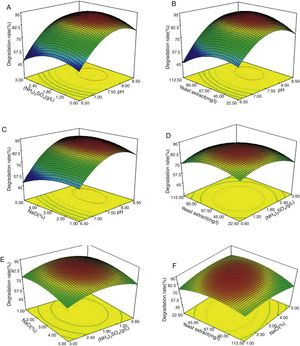
The effects of the independent factors (initial pH, concentrations of (NH4)2SO4, yeast extract and NaCl) and their interactions on testosterone degradation were examined. The 2D contour plot and 3D response surface were used to represent the regression equation graphically. Figure 4 presents the 3D response surface for the optimization of degradation and components of the medium.
The initial pH value of the medium played the most important role in testosterone degradation, which was consistent with the results of the Plackett–Burman design. Figure 4A–C clearly showed that the testosterone degradation rate changed most dramatically with the alteration of the initial pH. When the initial pH value of the medium changed from 6.5 to 7.5, the testosterone degradation rate increased from about 60% to 85%, respectivley. It proved that when pH is lower than 7.5, the medium is not suitable for strain N3 growth, or that a low pH could have a negative effect on some enzymes and proteins which are important to testosterone absorption and digestion. This result showed that pH has an important effect on the testosterone degradation rate; therefore, it should be considered as an imporatnat factor to isolate the testosterone-degrading strain.
The effect of other factors on testosterone degradationAs shown in Figure 4, when the concentration of one of the other three elements was changed, the degradation rate was also changed, but the effects were weaker than those of pH. When pH was kept at a constant level, the other factors did not have dramatic effects on testosterone degradation (Fig. 4A–F). For example, when pH was 7.5 (Fig. 4B) and the concentration of yeast extract changed from 22.5mg/l to 67.5mg/l, the degradation rate decrease was less than 10%. The maximum decrease in the degradation rate of testosterone was 10–15% with these three factor changes. (NH4)2SO4 and yeast extract are important nitrogen sources, and play an important role in the rates of chemical degradation17,18. In addition to supplying nitrogen, yeast extract also contains amino acids, peptides, vitamins, and several organic acids, which are necessary to organisms26–28. The main role of NaCl is to maintain the osmotic pressure of the cells2,17. A lower or higher concentration of NaCl could cause the cells to take up or lose water; either situation is harmful to cell survival and the ability to degrade chemicals.
Optimum conditions for the degradation of testosterone and optimized modelThe degradation rate reached the maximum value when the concentrations of (NH4)2SO4, yeast extract and NaCl were optimized. Overall, considering the main and interactive effects of the four factors, the optimal conditions were determined by solving the regression equation with the Design Expert 7.0 software. The final result is as follows: initial pH 7.95, 1.75g/l (NH4)2SO4, 68.06mg/l yeast extract, and 3.12% (w/v) NaCl.
For the purpose of testing this model for testosterone degradation, degradation experiments were carried out under optimal conditions. The result showed that 92% of testosterone (0.3mM) was degraded in 48h. Consistently, from the prediction of the RSM model, 95% of testosterone (0.3mM) was degraded in 72h. The results demonstrated that the model was accurate and the optimized conditions that we determined could be used for further research. The optimum concentration of 3.12% salt and pH 7.95 were similar to that of seawater of the South China Sea, which indicates that it could also show high testosterone degradation rate in the original environment. Since testosterone contamination has become a serious problem in the environment, strain N3 could be a potentially useful microorganism for testosterone pollution treatment.
ConclusionsIn this study, four testosterone-degrading strains N1, N3, N6 and N13 were isolated from the sediment in the marine fish culture area near Nanao Island, South China Sea. Strains N1, N3, N6 and N13 could grow in MM medium with 10mg/l testosterone as the sole carbon source. The capacity of the selected strains to degrade testosterone was further studied in two steps. Vibrio sp. N3 showed the highest testosterone degradation rate. Strain N3 could also degrade E2, EE2 and cholesterol. Based on the results of the Plackett–Burman design and response surface methodology with central composite design, the optimal conditions for testosterone degradation were the following: initial pH 7.95, shaker roll rate 150rpm, 25°C, 1.75g/l (NH4)2SO4, 0.25g/l K2HPO4, 68.06mg/l yeast extract, 0.25g/l glucose, and 3.12% (w/v) NaCl. Therefore strain N3 could be a potentially useful microorganism for steroid pollution treatment in the environment. Although this strain could degrade a wide range of substrates, several problems need to be addressed before bioremediation application. These issues include: the mechanism underpinning the degradation of these substrates, the enzyme(s) responsible for the breakdown of substrates, and the capacity to degrade the extremely low concentration of environmental steroids.
Conflict of interestThe authors of the paper have no financial and personal relationships with other people or organizations that could inappropriately influence (bias) this work.
This work was supported by National Nature Science Foundation of China (Nos. 31670117, 31770130, 41706143, 31870104) and Science & Technology Project of Guangdong Province, P.R. China (2014A020217017).