The objective is to describe the resistance mutation rate in protease and reverse transcriptase genes and sensitivity to different antiretrovirals in our environment.
MethodsWe performed an observational descriptive study in which we examined the samples provided at the Clinical Immunology Laboratory between April 2004 and April 2009. We analysed both the resistance tests and the sensitivity to different drugs in patients with therapeutic failure using Trugene HIV-1 Genotyping Kits®.
ResultsWe registered samples from 242 patients, 61 of which had no detectable resistance. The most prevalent mutations according to drug families were: for nucleoside analog reverse transcriptase inhibitors T215A/C/D/F/L/N/S/Y (24.10%), M184G/I/V/W (14.66%), M41J/L/R/T/W (11.24%) and K219E/G/H/N/R/T/W (10.24%). The highest levels of resistance corresponded to stavudine and lamivudine/emtricitabine, and tenofovir produced the least resistance in our environment. The non-analogues were K103N/R (23.98%), V179D/E/I/M/T (10.82%), A98E/G/S (10.53%) and K101E/P/Q/R (9.06%). Nevirapine presented greater resistance than efavirenz.
Protease inhibitors were L10F/I/V (15.95%), M36I/L (13.81%), A71I/T/V (13.10%) and 154L/S/V (7.38%). The darunavir/ritonavir combination was that which presented the least resistance, and tipranavir/ritonavir and lopinavir/ritonavir the most resistance.
ConclusionsAntiretroviral resistance and sensitivity to retroviral treatment in our environment was similar to results from other studies in Spain, but differed in the high level of resistance to lamivudine/emtricitabine and lopinavir/ritonavir.
El objetivo es describir la tasa de mutaciones de resistencia en los genes de la proteasa y la transcriptasa inversa y la sensibilidad de los diferentes antirretrovirales en nuestro medio.
MétodosEstudio observacional, descriptivo en el cual se estudiaron las muestras remitidas al laboratorio de Inmunología Clínica desde abril de 2004 hasta abril de 2009. Se analizaron tanto los test de resistencias, como el análisis de sensibilidad a los diferentes fármacos de pacientes en fracaso terapeutico mediante TRUGENE HIV-1 Genotyping Kit®.
ResultadosSe registraron las muestras de 242 pacientes, en 61 de ellos no se detectaron resistencias. Las mutaciones mas prevalentes según familia de fármacos fueron: para los inhibidores de la transcriptasa inversa análogos nucleosídicos T215A/C/D/F/L/N/S/Y (24,10%), M184G/I/V/W (14,66%), M41J/L/R/T/W (11,24%) y K219E/G/H/N/R/T/W (10,24%). La estavudina y la lamivudina/emtricitabina fueron los que mas resistencias presentaron, y el tenofovir es el que tiene menos resistencias en nuestro medio. En cuanto a los no análogos fueron K103N/R (23,98%), V179D/E/I/M/T (10,82%), A98E/G/S (10,53%) y K101E/P/Q/R (9,06%). Nevirapina presentó más resistencias que efavirenz.
Respecto a los inhibidores de la proteasa fueron L10F/I/V (15,95%), M36I/L (13,81%), A71I/T/V(13,10%) y I54L/S/V (7,38%). La combinación darunavir/ritonavir fue la que menos resistencias presentó junto con tipranavir/ritonavir, en contraposición lopinavir/ritonavir fue el que más resistencias obtuvo.
ConclusiónLa resistencia y sensibilidad al tratamiento antirretroviral en nuestro medio fue similar a la de otros estudios realizados en nuestro país, pero difiere y destaca un alto grado de resistencia a lamivudina/emtricitabina y lopinavir/ritonavir.
Acquired immunodeficiency syndrome (AIDS) has cost the lives of more than 25 million people since it was identified in 1981 and the effectiveness of pharmacology is multifactorial for these complex patients.1 Resistance to antiretroviral drugs is a major factor in HIV positive patients, and knowledge of these drugs is necessary to understand the evolution and result of treatments.2 Resistance analysis through genotype and phenotype testing has become widespread in recent years. The information these tests provide is a key tool in the hospital when establishing the most appropriate treatment, allowing greater effectiveness.3,4 Thus resistance testing is currently considered a basic procedure in the therapeutic monitoring of HIV positive patients, and is included in reference treatment guidelines as a highly useful procedure.5–7
Resistance may be transmitted to others, and may vary according to region, patient group and method used. The figures for recent infection by resistant viruses vary between 7.7% and 19.2%.8
Knowledge of factors that determine the development of resistances to various antiretroviral drugs may be useful in understanding treatment failure, and may help in planning therapeutic guidelines, which should be based on the results for each local situation.9
The aim of this study is to report the rate of resistance mutations of HIV-1 protease and reverse transcriptase genes and the sensitivity of various antiretrovirals in our environment, in order to understand the actual state of antiretroviral drug resistance in our hospital.
MethodsObservational, descriptive study analysing samples delivered to the Clinical Immunology laboratory from April 2004 to April 2009. The study population consisted of all patients who underwent resistance testing during the inclusion period, all of which experienced treatment failure. Plasma samples were separated within 3h from extraction and preserved in aliquots at −85°C. Viral load was determined by Branched DNA with the VERSANT 440 Molecular System® (Siemens Medical Solutions Diagnostics, Tarrytown, NY, USA), according to the manufacturer's instructions. The resistance testing was performed through Trugene HIV-1 Genotyping Kit® and the DNA Opengene® sequencing system (Siemens Healthcare diagnostics, Deerfield, USA) for detecting mutations in HIV-1 protease and reverse transcriptase, according to the protocol authorised by the manufacturer. The RNA was previously isolated using the QIAamp Viral RNA mini kit® (Qiagen, Hilden, Germany).10
The results were uploaded to a Microsoft Office Access® 2007 database where they were processed, selecting only those mutations detected that were clinically significant. The final data was treated with the SPSS® statistical analysis program version 12.0.
ResultsSamples from 242 patients were recorded: 153 from males (63.22%) and 89 from females (36.78%), with a mean age of 43.7±9.8 years. All patients suffered virologic failure due to multifactorial causes: lack of treatment adherence or possible drug resistance. No clinically significant wild-type mutations or resistance to antiretroviral drugs was found in 61 patients (25.20%). One hundred and eighty-one patients (74.8%) had clinically relevant mutations in one of the two regions. Of these, 153 (63.2%) had some mutation in both regions and 28 (11.6%) had mutation in only one of the regions.
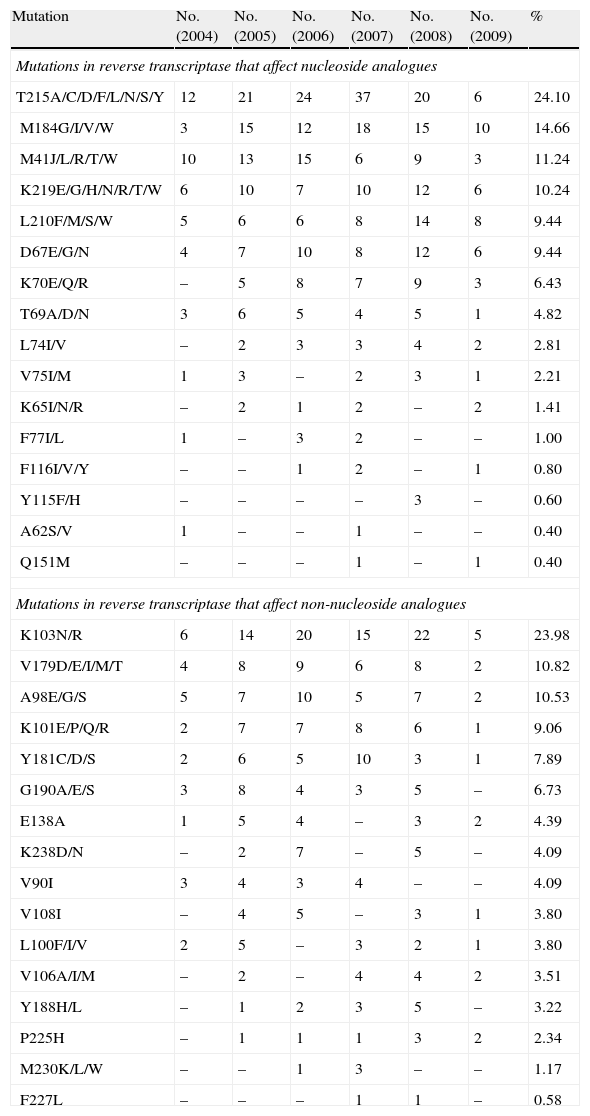
Tables 1 and 2 show the mutations detected in the reverse transcriptase and protease regions that affect antiretroviral drugs.
Mutations Detected in Various Regions of the Reverse Transcriptase That Affect Antiretroviral Drugs, From April 2004 to April 2009.
| Mutation | No. (2004) | No. (2005) | No. (2006) | No. (2007) | No. (2008) | No. (2009) | % |
| Mutations in reverse transcriptase that affect nucleoside analogues | |||||||
| T215A/C/D/F/L/N/S/Y | 12 | 21 | 24 | 37 | 20 | 6 | 24.10 |
| M184G/I/V/W | 3 | 15 | 12 | 18 | 15 | 10 | 14.66 |
| M41J/L/R/T/W | 10 | 13 | 15 | 6 | 9 | 3 | 11.24 |
| K219E/G/H/N/R/T/W | 6 | 10 | 7 | 10 | 12 | 6 | 10.24 |
| L210F/M/S/W | 5 | 6 | 6 | 8 | 14 | 8 | 9.44 |
| D67E/G/N | 4 | 7 | 10 | 8 | 12 | 6 | 9.44 |
| K70E/Q/R | – | 5 | 8 | 7 | 9 | 3 | 6.43 |
| T69A/D/N | 3 | 6 | 5 | 4 | 5 | 1 | 4.82 |
| L74I/V | – | 2 | 3 | 3 | 4 | 2 | 2.81 |
| V75I/M | 1 | 3 | – | 2 | 3 | 1 | 2.21 |
| K65I/N/R | – | 2 | 1 | 2 | – | 2 | 1.41 |
| F77I/L | 1 | – | 3 | 2 | – | – | 1.00 |
| F116I/V/Y | – | – | 1 | 2 | – | 1 | 0.80 |
| Y115F/H | – | – | – | – | 3 | – | 0.60 |
| A62S/V | 1 | – | – | 1 | – | – | 0.40 |
| Q151M | – | – | – | 1 | – | 1 | 0.40 |
| Mutations in reverse transcriptase that affect non-nucleoside analogues | |||||||
| K103N/R | 6 | 14 | 20 | 15 | 22 | 5 | 23.98 |
| V179D/E/I/M/T | 4 | 8 | 9 | 6 | 8 | 2 | 10.82 |
| A98E/G/S | 5 | 7 | 10 | 5 | 7 | 2 | 10.53 |
| K101E/P/Q/R | 2 | 7 | 7 | 8 | 6 | 1 | 9.06 |
| Y181C/D/S | 2 | 6 | 5 | 10 | 3 | 1 | 7.89 |
| G190A/E/S | 3 | 8 | 4 | 3 | 5 | – | 6.73 |
| E138A | 1 | 5 | 4 | – | 3 | 2 | 4.39 |
| K238D/N | – | 2 | 7 | – | 5 | – | 4.09 |
| V90I | 3 | 4 | 3 | 4 | – | – | 4.09 |
| V108I | – | 4 | 5 | – | 3 | 1 | 3.80 |
| L100F/I/V | 2 | 5 | – | 3 | 2 | 1 | 3.80 |
| V106A/I/M | – | 2 | – | 4 | 4 | 2 | 3.51 |
| Y188H/L | – | 1 | 2 | 3 | 5 | – | 3.22 |
| P225H | – | 1 | 1 | 1 | 3 | 2 | 2.34 |
| M230K/L/W | – | – | 1 | 3 | – | – | 1.17 |
| F227L | – | – | – | 1 | 1 | – | 0.58 |
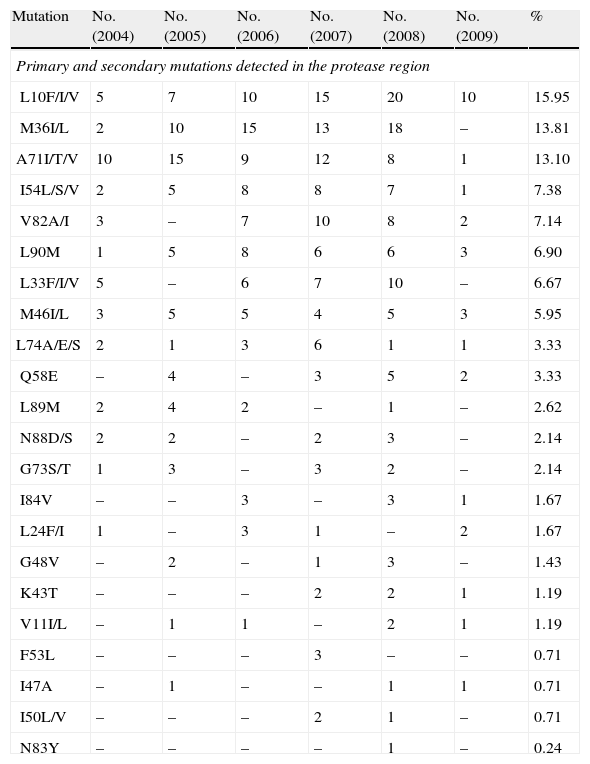
Mutations Detected in Protease That Affect Antiretroviral Drugs, From April 2004 to April 2009.
| Mutation | No. (2004) | No. (2005) | No. (2006) | No. (2007) | No. (2008) | No. (2009) | % |
| Primary and secondary mutations detected in the protease region | |||||||
| L10F/I/V | 5 | 7 | 10 | 15 | 20 | 10 | 15.95 |
| M36I/L | 2 | 10 | 15 | 13 | 18 | – | 13.81 |
| A71I/T/V | 10 | 15 | 9 | 12 | 8 | 1 | 13.10 |
| I54L/S/V | 2 | 5 | 8 | 8 | 7 | 1 | 7.38 |
| V82A/I | 3 | – | 7 | 10 | 8 | 2 | 7.14 |
| L90M | 1 | 5 | 8 | 6 | 6 | 3 | 6.90 |
| L33F/I/V | 5 | – | 6 | 7 | 10 | – | 6.67 |
| M46I/L | 3 | 5 | 5 | 4 | 5 | 3 | 5.95 |
| L74A/E/S | 2 | 1 | 3 | 6 | 1 | 1 | 3.33 |
| Q58E | – | 4 | – | 3 | 5 | 2 | 3.33 |
| L89M | 2 | 4 | 2 | – | 1 | – | 2.62 |
| N88D/S | 2 | 2 | – | 2 | 3 | – | 2.14 |
| G73S/T | 1 | 3 | – | 3 | 2 | – | 2.14 |
| I84V | – | – | 3 | – | 3 | 1 | 1.67 |
| L24F/I | 1 | – | 3 | 1 | – | 2 | 1.67 |
| G48V | – | 2 | – | 1 | 3 | – | 1.43 |
| K43T | – | – | – | 2 | 2 | 1 | 1.19 |
| V11I/L | – | 1 | 1 | – | 2 | 1 | 1.19 |
| F53L | – | – | – | 3 | – | – | 0.71 |
| I47A | – | 1 | – | – | 1 | 1 | 0.71 |
| I50L/V | – | – | – | 2 | 1 | – | 0.71 |
| N83Y | – | – | – | – | 1 | – | 0.24 |
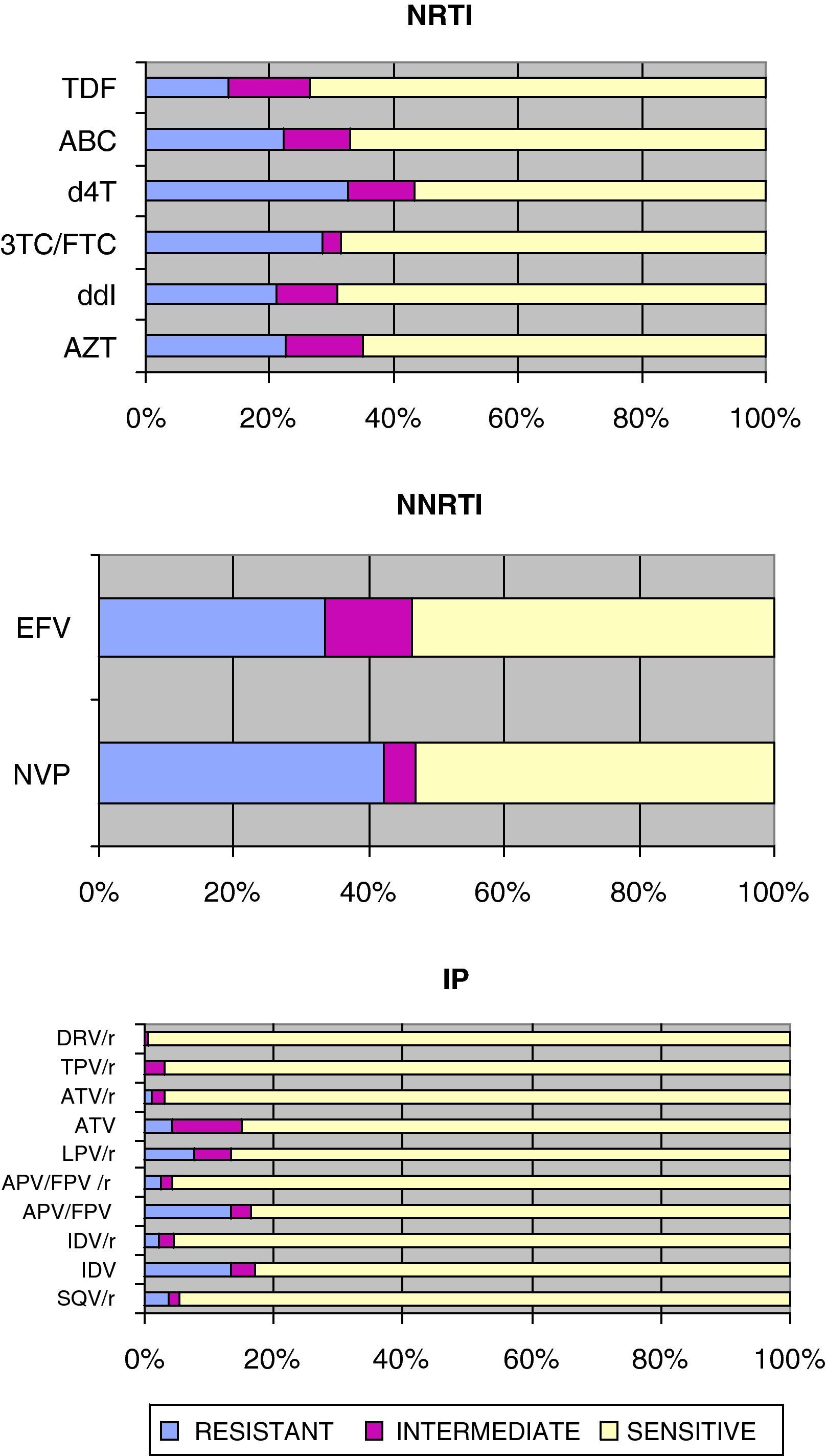
Fig. 1 shows the sensitivity of HIV-1 to the various families of antiretroviral drugs. NRTIs, d4T and 3TC/FTC had the most resistance, as opposed to TDF that had the least number of resistances in our environment.
Sensitivity of HIV-1 to reverse transcriptase and protease inhibitors:
3TC/FTC: lamivudine/emtricitabine; ABC: abacavir; APV/FPV: amprenavir/fosamprenavir; APV/FPV/r: amprenavir/fosamprenavir boosted with ritonavir; ATV: atazanavir; ATV/r: atazanavir/ritonavir; AZT: zidovudine; d4T: stavudine; ddI: didanosine; DRV/r: darunavir/ritonavir; EFV: efavirenz; IDV: indinavir; IDV/r: indinavir/ritonavir; LPV/r: lopinavir/ritonavir; NVP: nevirapine; SQV/r: saquinavir/ritonavir TDF: tenofovir; TPV/r: tipranavir/ritonavir.
Regarding NNRTIs, NVP had more resistance than EFV1, although both were sensitive to a similar percentage due to the higher percentage of intermediate resistance for EFV.
For the PIs, the DRV/r combination had the least number of resistances along with TPV/r, as opposed to unboosted PIs that had the most resistances. Among the boosted PIs, LPV/r had the most resistances.
DiscussionThe patients studied met current recommendations and therefore underwent resistance tests before changing treatment after virologic failure. In recent years, however, there has been a tendency to perform this test in treatment-naïve patients with the aim of selecting the optimal first treatment regimen, resulting in a cost-effective option.5,11
The study found that, in 25.20% of patients, no significant resistances were detected and sensitivity to drugs was total, indicating that treatment failure may be due to undocumented lack of adherence. However, patient pharmaceutical exposure to antiretroviral treatment is not known and we should take into account that greater exposure leads to a greater likelihood of generating mutations. There was also no data on viraemia, which is closely related to genotypic resistance.12,13
In terms of mutations found in the study, the most prevalent mutations affecting NRTIs coincide with those shown in other studies.4,14 These include T215A/C/D/F/L/N/S/Y (24.10%), M184G/I/V/W (14.66%), M41J/L/R/T/W (11.24%) and K219E/G/H/N/R/T/W (10.24%). The T215F/Y mutation affects most NRTIs, which compromises the response to AZT by itself. T215C/D/S also affects AZT and d4T. The M184V/I mutation compromises the response to 3TC+FTC by itself, affecting ABC+ddI to a lesser degree. It is the first to appear when they are used as part of the treatment.15 This mutation has been associated more with the combination of AZT+3TC and less with TDF+FTC.16 This mutation also causes hypersusceptibility to AZT, d4T and TDF. The M41L mutation is more specific to d4T and TDF, while K219E/R/T/W is more specific to AZT and d4T. Also noteworthy is the L74I/V mutation (2.81%), which by itself compromises the response to ddI, while K65R (1.41%), does so with TDF.
Also of note are the resistances known as TAM (thymidine analogue mutations), which cause resistance to AZT, d4T and TDF.
As for NNRTIs, the prevalence of mutations in this group has increased significantly in recent years.17 The most frequent were K103N/R (23.98%), V179D/E/I/M/T (10.82%), A98E/G/S (10.53%) and K101E/P/Q/R (9.06%), which affect both NVP and EFV. The K103N mutation was the most frequent, as well as in the evaluated studies.9
The mutations that affected the PIs were similar to those of comparative studies, but unlike those where the most frequent were I54L/V and V82A, in our case, it was L10F/I/V. This mutation was associated with resistance to each of the PIs in the presence of other mutations. L10F/I/V behaves as a frequent polymorphism in resistance sites in non-B subtype variants, in the same manner as M36I. Furthermore, I54L/S/V is characteristic of most PIs. Also of note is the detection of mutations, which by itself compromises drug response, although they appear in a small proportion. These mutations include G48V for SQV, I50L for ATV, I50V for FPV and I47A for LPV, all boosted with ritonavir.
As for the sensitivity of various drugs, the relationship between emergence of resistant mutations and virologic failure is quite clear in the case of NRTIs and NNRTIs. However, there are controversial data regarding PIs, in which virologic failure have been confirmed without evidence of resistance.18 In this study, TDF was the most sensitive of the NRTIs, in part because it is the most recent one, coinciding with the Molina et al. study.9
With regard to NNRTIs, the resistance mutations of NVP and EFV cause cross-resistance with the other element of the family, and therefore there is little room for action if detected.19 However, the recent marketing of etravirine leaves the door open in this situation, at the expense of future results.
Regarding PIs, we should consider that data on unboosted resistance pertains to the first years of the study.7 The Molina et al. study agrees on the greater efficacy of DRV/r and TPV/r, which are the most recent PIs.9 However, the same is not true with LPV/r, which has a high degree of resistance in our environment compared to the rest of the boosted PIs. This is explained in part by the mutations in the M46, I54 and I84 positions, which reduce the effectiveness of LPV/r to less than 30%.20
In conclusion, the resistance and sensitivity to HAART in our environment was similar to those of other studies performed in Spain, but unlike these studies ours showed a high degree of resistance to 3TC/FTC and LPV/r. It is advisable to conduct a behavioural analysis of resistance to antiretroviral drugs in order to understand the reality of this issue in our environment. From a multidisciplinary perspective, the data may be very useful in deciding the most effective therapy.
Conflict of InterestThe authors declare that they have no conflict of interest.
Please cite this article as: Fernández Lisón LC, et al. Tasa de mutaciones genotípicas y resistencia a antirretrovirales en un hospital general. Farm Hosp. 2011;35(4):191–196.