This study aimed to explore the underlying anti-asthma pharmacological mechanisms of conciliatory anti-allergic decoction (CAD) with a network pharmacology approach.
MethodsTraditional Chinese medicine related databases were utilized to screen the active ingredients of CAD. Targets of CAD for asthma treatment were also identified based on related databases. The protein-protein interaction network, biological function and KEGG pathway enrichment analysis, and molecular docking of the targets were performed. Furthermore, an asthma mouse model experiment involving HE staining, AB-PAS staining, and ELISA was also performed to assess the anti-asthma effect of CAD.
ResultsThere were 77 active ingredients in CAD, including quercetin, kaempferol, stigmasterol, luteolin, cryptotanshinone, beta-sitosterol, acacetin, naringenin, baicalin, and 48 related targets for asthma treatment, mainly including TNF, IL4, IL5, IL10, IL13 and IFN-γ, were identified with ideal molecular docking binding scores by network pharmacology analysis. KEGG pathway analysis revealed that these targets were directly involved in the asthma pathway, Th1 and Th2 cell differentiation, and signaling pathways correlated with asthma (NF-κB, IL17, T cell receptor, TNF, JAK-STAT signaling pathways, etc.). Animal experiments also confirmed that CAD could attenuate inflammatory cell invasion, goblet cell hyperplasia and mucus secretion. The levels of the major targets TNF-α, IL4, IL5, and IL13 can also be regulated by CAD in an asthma mouse model.
ConclusionThe anti-asthma mechanism of CAD possibly stemmed from the active ingredients targeting asthma-related targets, which are involved in the asthma pathway and signaling pathways to exhibit therapeutic effects.
Asthma is a common chronic respiratory disease especially endemic among children. According to the epidemiological survey by the World Health Organization in 2015, there were approximately 334 million people (4.9% of the world's population) suffering from asthma, and 250,000 people die of asthma prematurely each year.1 Recent years have witnessed a rising tendency of the morbidity and mortality of asthma worldwide, which poses a serious threat to human health and medical resources.2,3 Studies have shown that many risk factors, including genetic susceptibility, allergens, air pollution, climate change, and respiratory virus infection are closely related to asthma.4 The pathogenesis of asthma is complex and has not been clearly elucidated, but the airway inflammation, accompanied by airway hyperresponsiveness and airway remodeling, are well-acknowledged characteristics of asthma. In the allergic reaction of asthma, allergens or pathogens that enter through body surfaces (e.g., the skin or lungs) are phagocytized by antigen-presenting cells, such as dendritic cells, which come to mature with the help of different cytokines.5 Th2 cells can produce pro-inflammatory cytokines, including IL4, IL5, and IL13 to exert their effects on many other cell types, including B cells, eosinophils, mast cells, epithelial cells, and airway goblet cells, and can also induce immunoglobulin E (IgE) production by B cells, resulting in the regulation of inflammation in asthma. Some drugs such as glucocorticoids, β2 receptor agonists, anti-cholinergic drugs, theophylline and leukotriene receptor antagonists, have also been approved for asthma treatment; however, the side effects and acquired resistance limit their clinical application.6–8 Thus, developing more safe and effective drugs for asthma therapy remains urgent and crucial.
Over thousands of years, traditional Chinese medicine has provided excellent therapeutic effects for asthma treatment in clinical practice.9 Traditional Chinese medicine possesses the advantages of “simple, convenient, economic, effective and individualized therapy”, which is in good accordance with the concept of modern medicine.10 Conciliatory anti-allergic decoction (CAD), a modified decoction comprising 10 herbs originating from the classical formula Xiao Chai Hu Tang by the well-known Chinese physician Zhang Zhongjing, has been widely used for asthma treatment in hospitals. Our former clinical research revealed that CAD can effectively reduce the frequency of asthma attacks and respiratory tract infection in children with asthma, enhance the immune function and anti-allergic ability of respiratory mucosa, and alleviate airway inflammation.11 However, the therapeutic mechanism of CAD in asthma still remains unclear.
Our current study was designed to explore the anti-asthma mechanisms of CAD. Network pharmacology study was applied in this work since it can provide a novel strategy to uncover the bioactive ingredients and underlying mechanisms of CAD from a systemic and holistic perspective.12 With the approach of network pharmacology, the active ingredients and related targets relevant to asthma were screened out, an ingredients-targets network and a protein–protein interaction (PPI) network were constructed, gene functional enrichment analysis, and molecular docking of the targets were conducted to clarify the pharmacological mechanisms of the targets and ingredients in CAD. Furthermore, animal experiments were also performed to validate the pharmacological mechanisms of CAD in an asthma mouse model.
Materials and methodsScreening of the active ingredients of CADThe CAD is composed of ten traditional Chinese herbs, including Radix Bupleuri (Chinese pinyin name Chaihu), Scutellariae Radix (Chinese pinyin name Huangqin), Pseudostellariae Radix (Chinese pinyin name Taizishen), Arum Ternatum Thunb. (Chinese pinyin name Banxia), Radix Salviae (Chinese pinyin name Danshen), Ephedra Herba (Chinese pinyin name Mahuang), Fritillariae Thunbrgii Bulbus (Chinese pinyin name Zhebeimu), Farfarae Flos (Chinese pinyin name Kuandonghua), Cicadae Periostracum (Chinese pinyin name Chantui), and Licorice (Chinese pinyin name Gancao). The ingredient information of these ten herbs was searched from the TCMSP database (http://ibts.hkbu.edu.hk/LSP/tcmsp.php). Data on the molecule name, 2D structure, Pubchem ID, pharmacological and molecular properties of the ingredients could all be obtained from that database. The ADME system parameters were used as the criteria to select the candidate active ingredients in each herb with oral bioavailability (OB) ≥30, drug-likeness (DL) ≥0.18 and half-life (HL) ≥4 based on the suggestion by the TCMSP database.
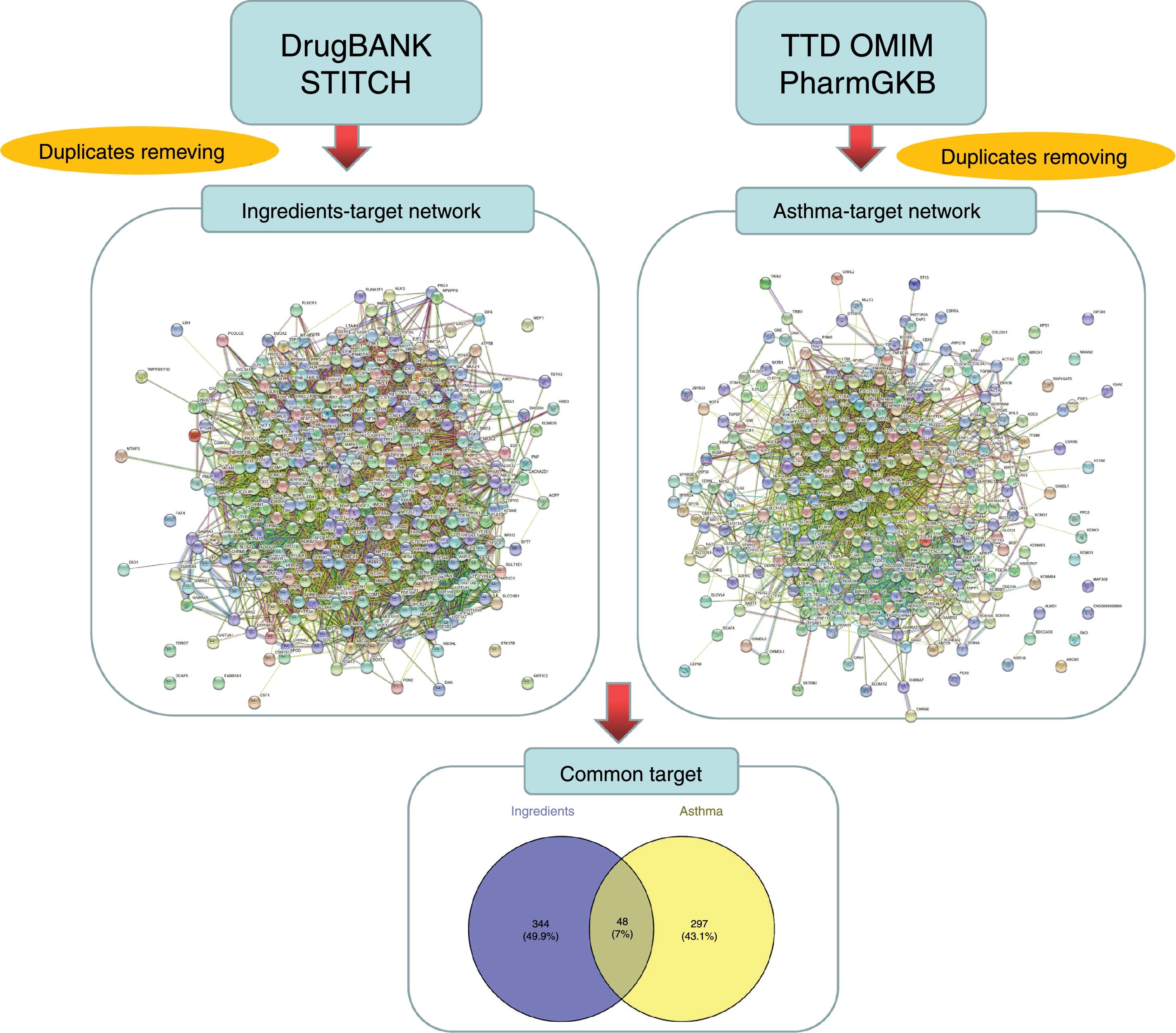
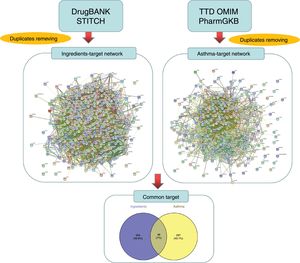
Prediction and screening of candidate targetsThe targets linked to the active ingredients in CAD were searched and predicted from the TCMSP database and STITCH database (http://stitch.embl.de/, ver. 5.0) with the ‘Homo sapiens’ species setting. The targets related to asthma were identified from the TTD (http://bidd.nus.edu.sg/group/cjttd/), OMIM (http://www.omim.org/), and PharmGKB (https://www.pharmgkb.org/) databases with ‘asthma’ as the input keyword. The target gene information, including IDs and names, was confirmed and standardized using UniProt (http://www.uniprot.org/) and duplicate targets were removed to obtain drug-related targets and disease-related targets, respectively. The Venny diagram online tool (http://bioinfogp.cnb.csic.es/tools/venny/) was employed to screen the overlapped targets between ingredients and diseases.
Protein–protein interaction (PPI) and gene functional enrichment analysis of the targetsThe STRING database (https://string-db.org/) was used to explore the interactions, and the hub gene targets among the candidate targets were screened with the ‘Homo sapiens’ species setting. The interaction data obtained from STRING were imported into Cytoscape software to construct the PPI network, and the topological properties of the network were analyzed with the plugin tool “Network analyzer”. For the candidate targets, the Cytoscape software plugin tool Clue GO was applied to perform the biological process and KEGG pathway enrichment analysis with terms, and P value <0.05 was set as the significance criterion for the terms.
Molecular dockingThe crystal structure of the target protein was obtained from the Protein Data Bank (http://www.rcsb.org/), and the 2D structure of the compound was downloaded from the PubChem database (https://pubchem.ncbi.nlm.nih.gov/). The docking exercise was conducted with the systemDock online tool (http://systemsdock.unit.oist.jp/iddp/home/index).
Network constructionThe ingredient-target network and target-related pathway network were constructed by the Cytoscape software and analyzed with the plugin tool “Network analyzer” to obtain the topological properties of the network.
Pharmacological verificationAnimalsBALB/c mice weighting 18–20g were purchased from the Centre of Experimental Animals at the Shanghai SLAC Laboratory Animal Co. Ltd. (Shanghai, China). The animal experiments were approved by the Ethics Committee of Zhejiang Traditional Chinese Medicine University (Hangzhou, China), and all animal procedures were carried out in accordance with the National Institutes of Health Guide for the Care and Use of Laboratory Animals. The mice were housed for a week at standard room temperature (20±2°C) and relative humidity (55±10%) under a 12h light/dark cycle, with free access to water and food.
Experimental design and drug administrationA total of 60 BALB/c mice were randomly divided into six groups (n=10) as follows: control group; ovalbumin (OVA) group; OVA+low CAD group (10mg/kg); OVA+medium CAD group (20mg/kg); OVA+high CAD group (40mg/kg); and OVA+dexamethasone (Dex) group (0.5mg/kg). The OVA-induced asthma mouse model was established as previously published with slight modifications.13 Briefly, the mice were sensitized by intraperitoneal injection of 2mg/ml OVA/Al(OH)3 gel in a total volume of 0.5ml or saline in the control group on days 1 and intraperitoneal injection of 0.2ml on day 13 to enhance sensitization. From day 23, the mice were then challenged by intranasal inhalations with OVA (10mg/mL) or PBS aerosol challenges for 30min three times a week for 8 weeks. CAD (10, 20, 40mg/kg) or Dex was administered intragastrically 1h prior to OVA challenge. The control and OVA groups received PBS on the same schedule with the same volume.
ELISA assayTwenty-four hours after the last OVA challenge, the mice were anesthetized with an inhalation of diethyl ether and sacrificed by exsanguination. Bronchoalveolar lavage fluid (BALF) was obtained by intratracheal instillation, and the lungs were lavaged three times with 0.8ml of sterile PBS. The BALF from each sample was centrifuged, and supernatants were stored at −80°C for subsequent analysis. The levels of TNF-α, IL4, IL5, IL10, and IL13 in BALF were measured by ELISA kits according to the manufacturer's instructions.
HE staining and AB-PAS stainingLung tissues from each group were collected and fixed in 10% buffered formalin for 24h, dehydrated, embedded in paraffin, and then cut into approximately 3-μm tissue sections. Subsequently, the tissue sections were stained with hematoxylin and eosin (H&E) to evaluate the degree of peribronchial and perivascular inflammation, and stained with Alcian blue-periodic acid Schiff (AB-PAS) to identify goblet cells in the epithelium and measure mucus production, respectively. The degree of peribronchial inflammation was scored in a blinded manner according to the following criteria: 0, no cells; 1, a few cells; 2, a ring of cells one cell layer deep; 3, a ring of cells two to four cells deep; and 4, a ring of cells of more than four cells deep. The degree of mucus production and goblet cell hyperplasia in the airway epithelium were also quantified in a blinded manner using a five-point scoring system: 0, no goblet cells; 1, 25%; 2, 25–50%; 3, 50–75%; and 4, 75%.14
Statistical analysisThe experimental data are presented as the mean±standard deviation (SD). Differences between groups were analyzed using one-way analysis of variance (ANOVA) followed by Bonferroni's test. P<0.05 was considered to be significant.
ResultsActive ingredients and candidate targets in CADA total of 77 active ingredients were screened from the TCMSP database according to the ADME criteria, with six compounds in Radix Bupleuri, 13 compounds in Scutellariae Radix, three compounds in Pseudostellariae Radix, five compounds in Arum Ternatum Thunb., 27 compounds Radix Salviae, nine compounds in Ephedra Herba, one compound in Fritillariae Thunbrgii Bulbus, four compounds in Farfarae Flos, two compounds in Cicadae Periostracum, and 27 compounds in Licorice. Although the ingredients in CAD are complex, some compounds were repeated among the herbs, like quercetin, kaempferol, stigmasterol, beta-sitosterol, luteolin, acacetin, naringenin, and baicalin, which exist in more than two herbs of CAD. Based on the DrugBank and STITCH databases, a total of 392 proteins were linked to the 77 identified ingredients, thus forming an ingredient-related target network (see Fig. 1). Based on the TTD, OMIM, and PharmGKB databases, a total of 345 proteins were identified to be related to asthma in the asthma target network. According to the Venny diagram, 48 proteins were found to be overlapped in the two networks and thus concluded as CAD ingredient-related targets for asthma treatment.
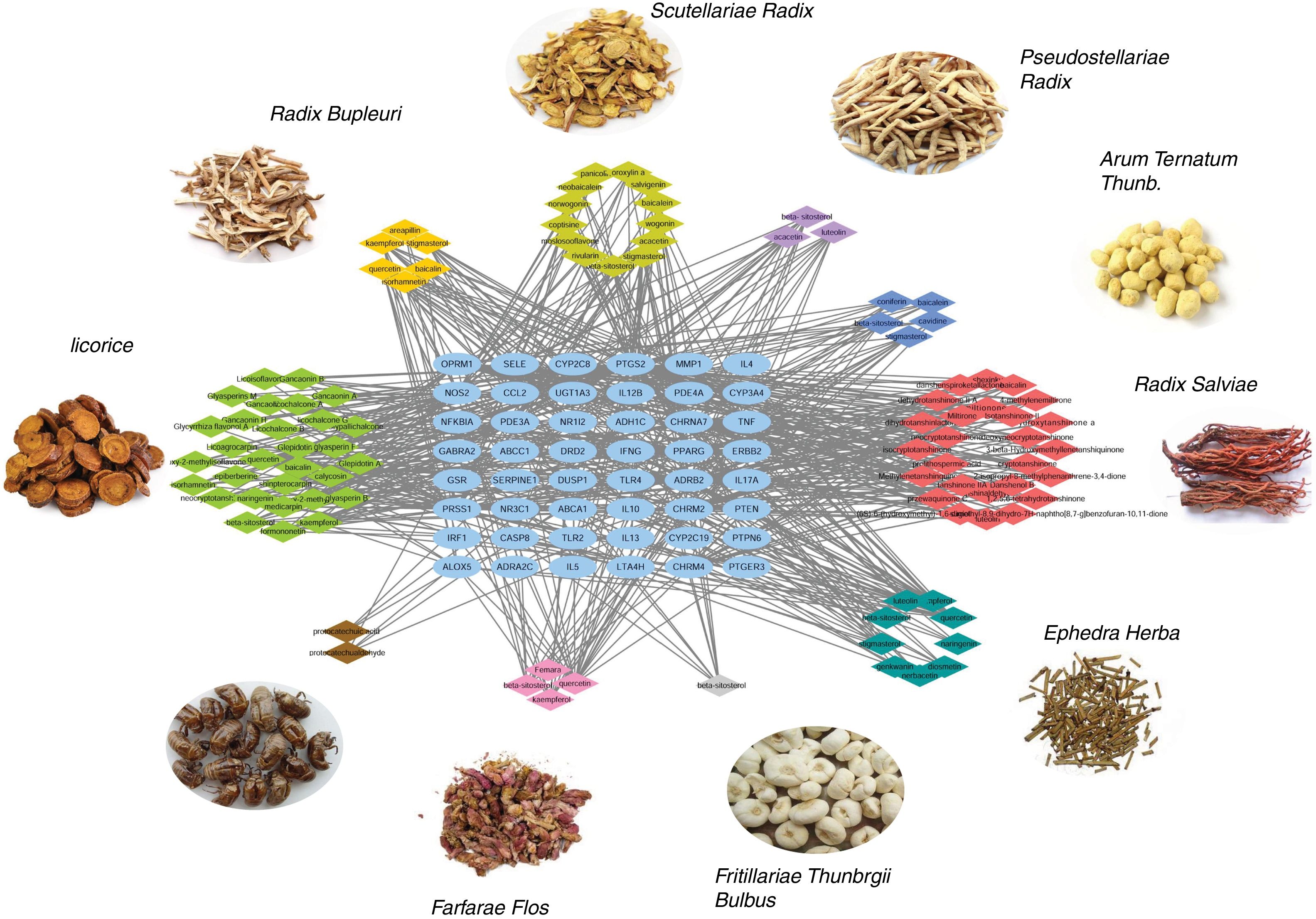
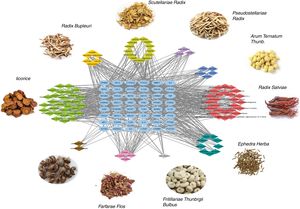
Ingredient-target network of CAD for asthma treatmentThe ingredient-target network of CAD for asthma treatment was visualized by Cytoscape and is shown in Fig. 2. There were 147 nodes and 666 edges in the network. 48 blue elliptic nodes represent 48 targets, and diamond nodes with different kinds of colors represent ingredients in different herbs. The six compounds in Radix Bupleuri targeted 33 proteins in the network, 13 compounds in Scutellariae Radix targeted 26 proteins in the network, three compounds in Pseudostellariae Radix targeted 21 proteins in the network, five compounds in Arum Ternatum Thunb. targeted 23 proteins in the network, 27 compounds in Radix Salviae targeted 29 proteins in the network, nine compounds in Ephedra Herba targeted 33 proteins in the network, one compound in Fritillariae Thunbrgii Bulbus targeted nine proteins in the network, four compounds in Farfarae Flos targeted 28 proteins in the network, two compounds in Cicadae Periostracum targeted six proteins in the network, and 27 compounds in Licorice targeted 33 proteins in the network. Quercetin had the highest degree with 20 proteins targeted, followed by kaempferol with 12 targets, wogonin with 10 targets, stigmasterol, luteolin, beta-sitosterol and acacetin with nine targets, etc. The ingredient-target network has properties of complex ingredients, multiple targets, and close interactions between ingredients and targets, thus forming a network for asthma treatment.
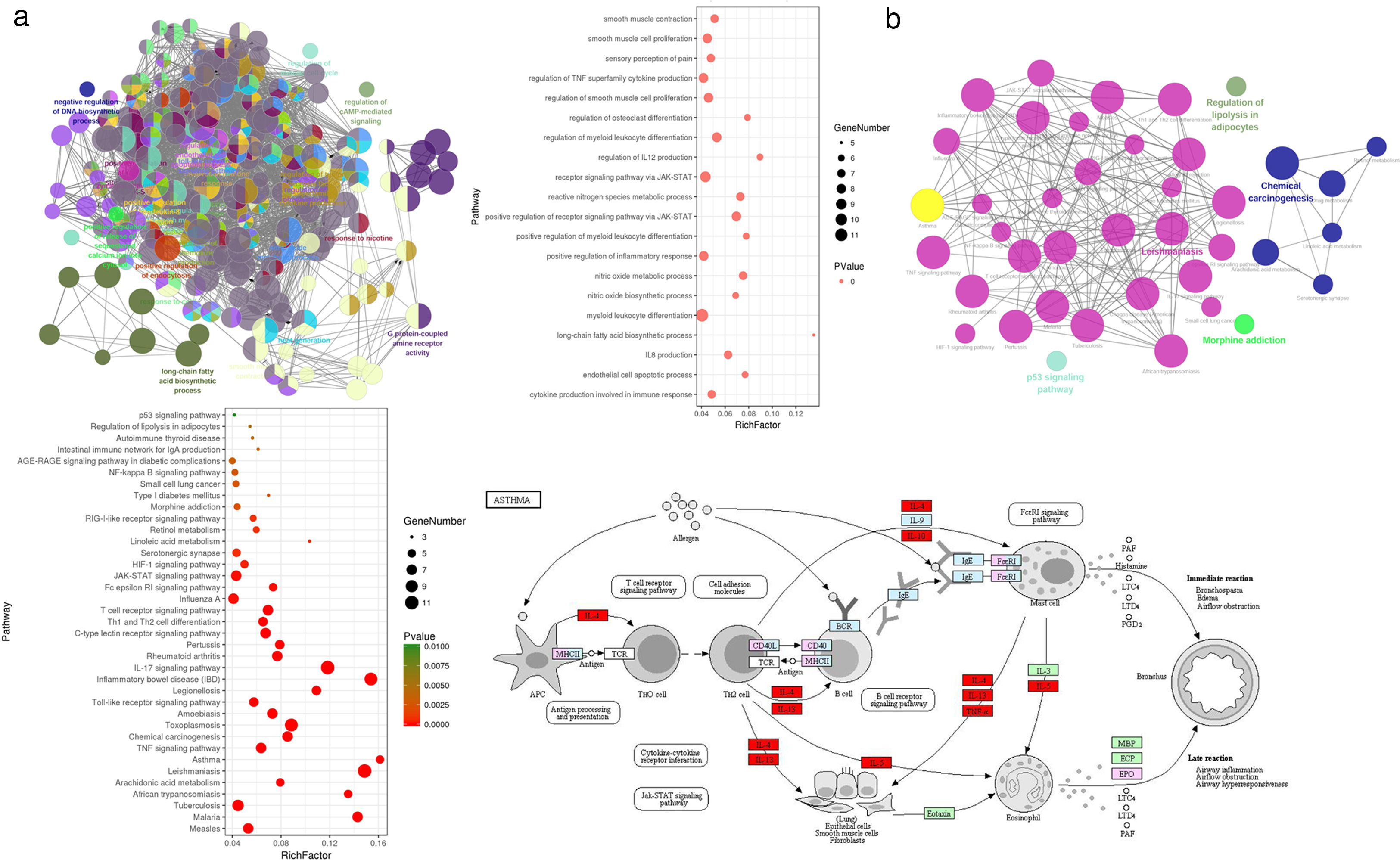
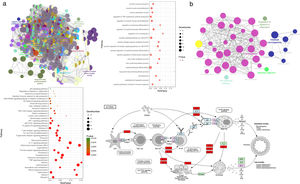
Gene Ontology enrichment analysisWith the Cytoscape software plugin tool Clue GO, gene functional enrichment analysis was performed (see Fig. 3a). Each node represents a biological process term, and node size represents term enrichment significance. Different colors represent different clusters. Gene biological process analysis showed that these 48 targets were enriched in 225 significant terms of biological process, and these biological process terms also interact closely with each other and are assigned in several clusters, thus forming a complex and compact network. The top 20 terms with their respective P values are also presented. These targets were mainly enriched in biological processes including cytokine production involved in immune response, the positive regulation of inflammatory response, regulation of IL12 production, IL8 production, the positive regulation of receptor signaling pathway via JAK-STAT, the regulation of TNF superfamily cytokine production, endothelial cell apoptotic process, nitric oxide metabolic/biosynthetic process, etc.
Gene functional enrichment analysis of the targets. (a) Gene Ontology enrichment analysis. In Gene Ontology enrichment network, each node represents a biological process term, with different colors assigned to different clusters. The top 20 significant terms of biological processes were also presented as a bubble diagram. (b) Target-related KEGG pathway analysis. KEGG pathway network and the enriched pathways bubble diagram of the targets were presented; these targets were enriched in the asthma pathway, and the genes with red color in the asthma pathway represent CAD related targets. The figure of the asthma pathway was downloaded from the KEGG database (https://www.genome.jp/kegg/).
KEGG pathway analysis of the targets was also conducted, and the results are shown in Fig. 3b. KEGG pathway network showed that these targets were significantly enriched in 40 pathway terms, which were assigned into five clusters and interacted closely with each other. The significant pathways that these targets were enriched in mainly include the asthma pathway, IL17 signaling pathway, T cell receptor signaling pathway, TNF signaling pathway, Th1 and Th2 cell differentiation, JAK-STAT signaling pathway, HIF-1 signaling pathway, NF-κB signaling pathway, etc. The pathway analysis results also demonstrated that each target was involved in several pathways, and accordingly, each pathway was enriched with several targets, thus forming a multiple target–multiple pathway network that directly or indirectly affects the occurrence and progression of asthma. Among these pathways, the asthma pathway was directly related to asthma, and five out of the 48 targets (TNF, IL4, IL5, IL10, IL13) were involved in this pathway with a term P value=1.16E−06 (Fig. 3b), indicating that CAD might interact with these targets, thus directly influencing the asthma pathway to play a protective role for asthma treatment.
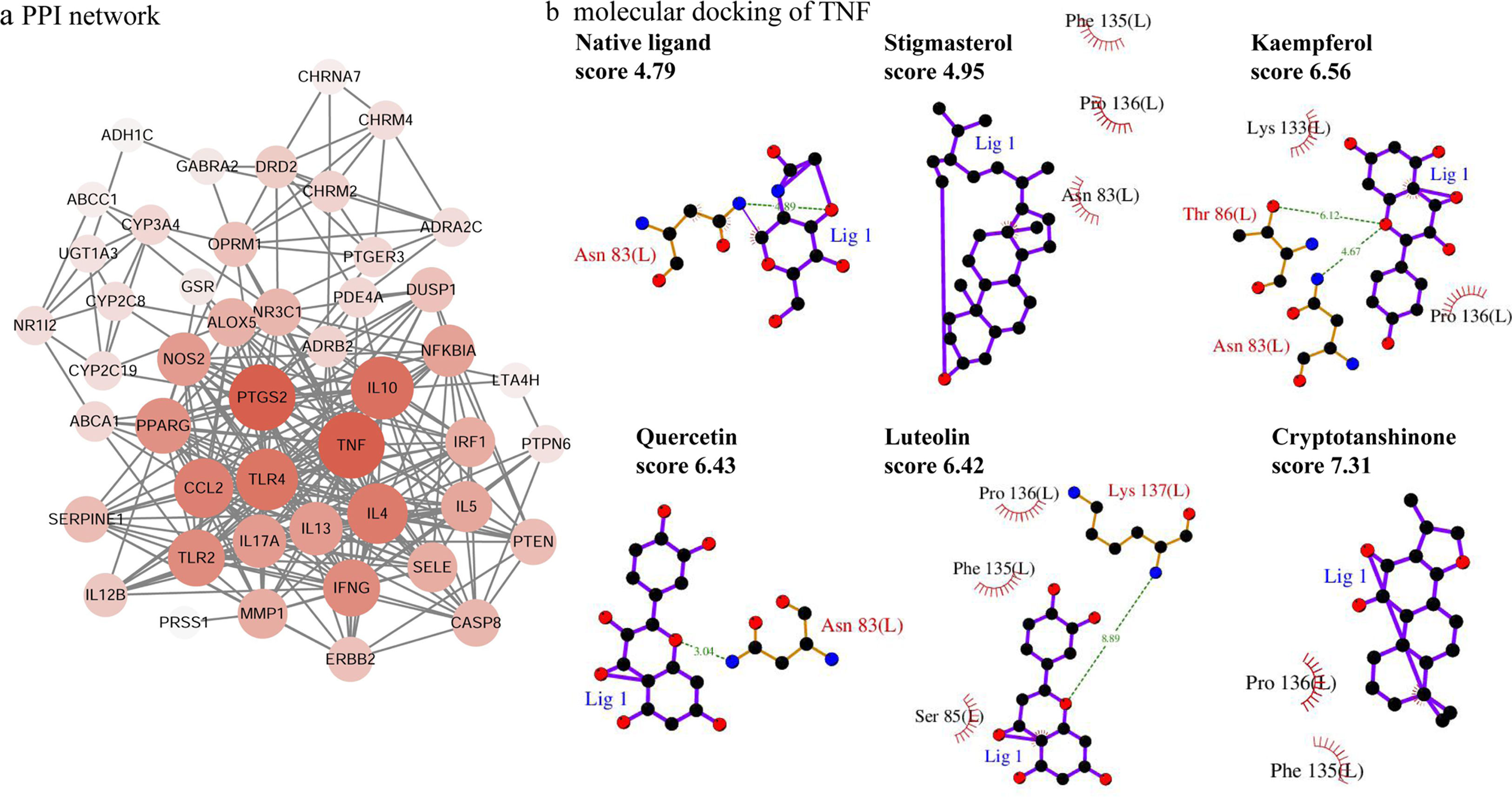
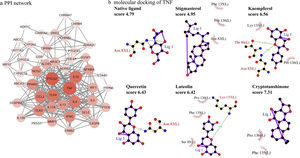
PPI network analysisThe PPI network is presented in Fig. 4a, there were 48 nodes and 285 edges in the network with a medium degree of six for each node. The network topological properties of the targets, such as degree, and closeness centrality were also analyzed according to the “Network analyzer” tool in Cytoscape. The node size and color of each target were positively related to the node degree. TNF (degree=29), PTGS2 (degree=29), IL10 (degree=26), TLR4 (degree=25), IL4 (degree=24), CCL2 (degree=23), IFNG (degree=21), and TLR2 (degree=21) were selected as the important targets from the PPI network with degree >20.
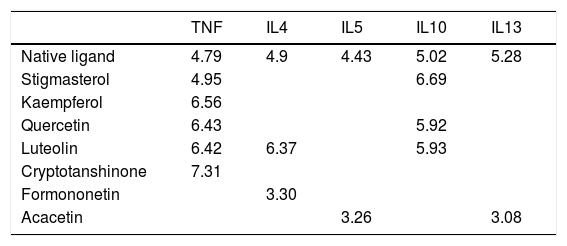
Molecular docking of the ingredients binding to asthma pathway related targetsSince the KEGG pathway analysis revealed that CAD's ingredient-related targets were involved in the asthma pathway, we used molecular docking analysis to validate the binding property of the asthma-related targets TNF, IL4, IL5, IL10, IL13 and the linked active ingredients with docking score. As shown in Fig. 4b, TNF native ligand had a docking score of 4.79, while stigmasterol, kaempferol, quercetin, luteolin, and cryptotanshinone in CAD that targeted TNF all had higher scores than that of the native ligand, which suggested that these active ingredients possess ideal interactions with TNF. The docking results of all five targets are shown in Table 1. It can be concluded that the majority of the ingredients have a higher score than the native ligand for the corresponding targets, which indicates that the ingredients in CAD can closely interact with the predicted targets and can thus influence asthma-related pathways.
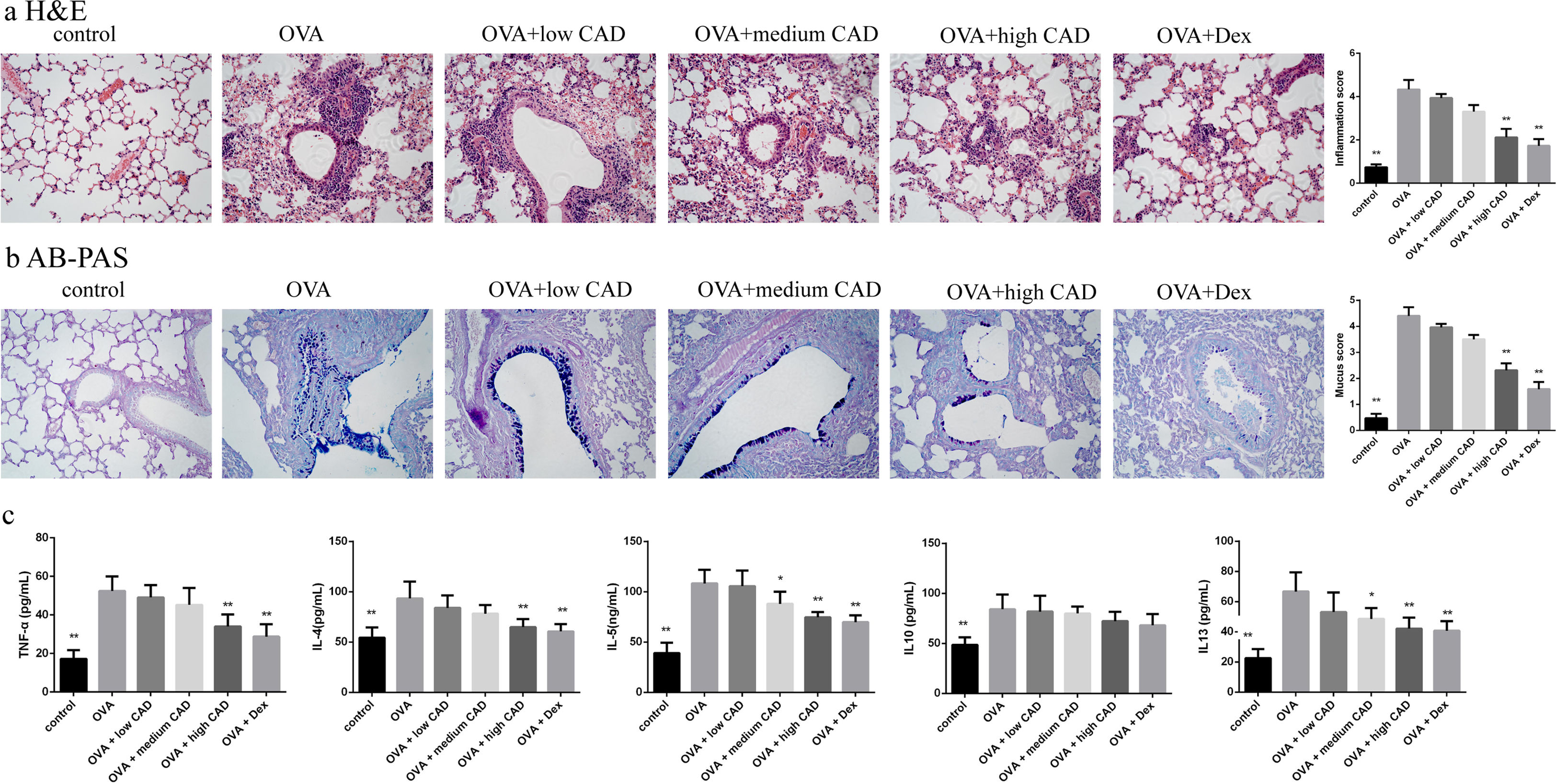
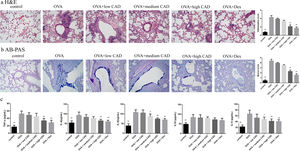
Effects of CAD on the histological changes in OVA-induced asthmatic mice lungsTo evaluate the therapeutic effect of CAD, histological studies of the lung tissues in different groups were performed. Compared with the control group, it can be observed that there was abundant inflammatory cell invasion into the peribronchial and perivascular areas in OVA-induced asthma mouse lung tissues stained by HE (see Fig. 5a). When CAD or Dex was administered at different doses, the inflammatory cell invasion was markedly attenuated, especially in the high CAD group and Dex group (P<0.05). AB-PAS staining was used to evaluate the presence of goblet cell hyperplasia and mucus secretion. As shown in Fig. 5b, the overexpression of goblet cell hyperplasia and mucus oversecretion can be observed in the bronchial airways of OVA group lung tissues. CAD in high doses (40mg/kg) and Dex can efficiently alleviate goblet cell hyperplasia and mucus secretion compared with the asthma model group (P<0.05).
Effect of CAD on OVA-induced asthmatic mice. (a) Effect of CAD on airway inflammation of asthma mice (HE staining, ×400); (b) effect of CAD on mucus hypersecretion of asthma mice (AB-PAS staining, ×400); (c) effect of CAD on TNF-α, IL4, IL5, IL10, and IL13 levels in mice BALF. The data presented are the means±SD, *P<0.05 or **P<0.01 vs. the OVA group. BALF: bronchoalveolar lavage fluid.
The levels of TNF-α, IL4, IL5, IL10, and IL13 in BALF were detected with ELISA and the results are shown in Fig. 5c. Compared to that in the normal control group, the levels of TNF-α, IL4, IL5, IL10, and IL13 in the OVA-induced asthma group were significantly elevated (P<0.01). After administration of CAD at different doses, the levels of these pro-inflammatory cytokines decreased to varying degrees. For TNF-α, CAD (40mg/kg) or Dex could significantly reduce the level of TNF-α in BALF (P<0.01), the same trend was also observed for IL4. For IL5 and IL13, the administration of 20mg/kg CAD reduced the levels in BALF (P<0.05), and with 40mg/kg CAD or Dex, the levels decreased significantly (P<0.01). For IL10, the changes in level among the experiment groups were not significant.
DiscussionAsthma is a chronic inflammatory disease of the airway with high morbidity and mortality globally. Numerous traditional Chinese medicines have excellent therapeutic effects for asthma, including CAD. Compared with the well-applied treatments in the clinic, the pharmacological mechanisms of CAD have not been researched clearly. In the present study, we applied network pharmacology to explore the anti-asthma mechanisms of CAD. A total of 77 active ingredients and 48 asthma-related targets of CAD were identified from the databases. Ingredient-target network and PPI network revealed that these ingredients and targets interact closely with each other. Pathway enrichment analysis also showed that these targets were directly or indirectly involved in asthma-related pathways. The molecular docking exercise showed that the majority of the active ingredients have a higher binding score than the native ligand binding to the corresponding targets. In the OVA-induced asthma mouse model, CAD administration efficiently attenuated airway inflammation and mucus production, and the expression of the hub targets were also decreased with CAD treatment.
CAD comprises 10 herbs, Radix Bupleuri, Scutellariae Radix, Pseudostellariae Radix, Arum Ternatum Thunb., Radix Salviae, Ephedra Herba, Fritillariae Thunbrgii Bulbus, Farfarae Flos, Cicadae Periostracum, and Licorice. Most of these herbs have been proven to have anti-inflammatory and anti-allergic activities, due to the complex active compounds in these herbs. Quercetin, one the most important ingredients, was found in four herbs of CAD with 20 targets; it was reported to regulate Th1/Th2 balance in asthma, reduce the level of IL4, and increase the level of IFN-γ, and restrain antigen-specific IgE antibody formation.15 Kaempferol, identified in four out of the ten herbs of CAD, was reported to suppress eosinophil infiltration and airway inflammation in airway epithelial cells and in mice with allergic asthma by disturbing NF-κB signaling.16 Baicalin, screened from Radix Bupleuri, Arum Ternatum Thunb., and Radix Salviae, was reported to inhibit airway remodeling in asthmatic mice by decreasing the expression of TGF-β1, IL13, and VEGF and inhibiting the activation of the extracellular signal-regulated kinase pathway.17 Stigmasterol in CAD was also reported with significant anti-asthmatic properties and had suppressive effects on the key features of allergen-induced asthma.18 Wogonin identified from Scutellariae Radix attenuated OVA-induced airway inflammation in a mouse model of asthma via the suppression of IL4/STAT6 signaling.19 These studies demonstrated that the major active ingredients identified from CAD were efficient for asthma treatment and accordingly confirmed the therapeutic effect of CAD.
The pathway enrichment analysis in the present study revealed that TNF, IL4, IL5, IL10, IL13 were directly enriched in the asthma pathway. And several targets including TNF, PTGS2, IL10, TLR4, IL4, CCL2, IFN-γ, and TLR2 were also regarded as important targets according to their topological properties in the PPI network. TNF-α and IFN-γ belong to Th1 cell cytokines, IL4, IL5, and IL13 belong to Th2 cell cytokines, and the imbalance of Th1/Th2 cells is related to the pathogenesis of asthma. TNF-α, the most studied pro-inflammatory cytokine of the TNF family, is produced by several pro-inflammatory cells (mainly macrophages, but also monocytes, dendritic cells, B-cells, CD4+ cells, neutrophils, mast cells and eosinophils) and is known to be crucial in the pathogenesis of asthma.20 Its elevated expression can be involved in the development and progression of airway pathology in asthma, and has recently been highlighted as potentially important for asthma. The development of neutralizing biological agents against TNF-α is also a therapeutic strategy for asthma with improvement in lung function, airway hyper-responsiveness and quality-of-life in patients.21 The administration of IFN-γ has the ability to elevate airway hyper-responsiveness via the up-regulation of neurokinin A/neurokinin-2 receptor signaling in severe asthma.22
The Th2 cytokines including IL4, IL5, and IL13 are critical in the pathogenesis of asthma, since they contribute to hallmarks of this disease, including airway inflammation, airway eosinophilia, increased mucus production, goblet cell hyperplasia, production of allergen-specific IgE and development of airway hyper-responsiveness.23 IL4 plays a key role in inducing T cell polarization into Th2 cells, and the subsequent generation of IL4, IL5, and IL13 by Th2 cells.5 IL5 can induce eosinophilia in lung tissues during asthma via the production of eotaxins to activate the recruitment of eosinophils to the lung tissues. Similar to IL4 and sharing the same signaling pathways, IL13 and its receptor IL13Rα1 can be detected in eosinophils, B cells, macrophages, smooth muscle cells, lung epithelial cells, airway goblet cells, and endothelial cells. Antagonists targeting at IL4, IL5, and IL13 have also been developed as a therapeutic strategy in asthma.24,25 The active ingredients, including quercetin, kaempferol, stigmasterol, luteolin, and cryptotanshinone in CAD, were identified to target TNF, IL4, IL5, IL10, and IL13, and the molecular docking also proved the ideal binding score between these cytokines and active ingredients. In the asthma mouse model of our study, elevated levels of these Th1 and Th2 cytokines were also detected, and the CAD-treated group showed a significant decrease in cytokines expression in a dose-dependent manner. It can be indicated that CAD efficiently targets Th1 and Th2 cell cytokines, maintains Th1/Th2 balance, and inhibits the asthma pathway, thus exerting curative effects on asthma.
In the pathogenesis of asthma, except for the central role of Th2 cells cytokines (IL4, IL5, IL10, IL13), or Th1 cells cytokines (TNF-α, IFN-γ), the signaling pathways involved with various targets are also vitally important. The NF-κB signaling pathway is one of the most important cellular signal transduction pathways that is essential for apoptosis, tumorigenesis, inflammation, viral infections, and various autoimmune diseases. The expression of the NF-κB signaling pathway can be abnormally activated in asthma, thus regulating many downstream targets and the secretion of pro-inflammatory cytokines to accelerate the progression of asthma.26 Studies have found that airway inflammation can be alleviated via inhibition of the NF-κB pathway in asthma.27,28 The differentiation of Th1 and Th2 cells from naive T cells occurs primarily via the JAK/STAT signaling pathway. IFN-γ-induced JAK/STAT signaling pathway is reported to be responsible for glucocorticoid insensitive in airway epithelial cells, and this steroid responsiveness of insensitive can be restored by the transfection of cells with siRNA-STAT1.29
In conclusion, the protective effect and underlying mechanism of CAD on asthma was explored in the current study. The multiple active ingredients in CAD, including quercetin, kaempferol, stigmasterol, luteolin, cryptotanshinone, beta-sitosterol, acacetin, naringenin, baicalin and related targets for asthma, mainly including TNF, IL4, IL5, IL10, IL13, and IFN-γ, were identified with ideal binding scores by network pharmacology. KEGG pathway analysis revealed that these targets were involved in the asthma pathway, Th1 and Th2 cell differentiation, and signaling pathways correlated with asthma (NF-κB signaling pathway, IL17 signaling pathway, T cell receptor signaling pathway, TNF signaling pathway, JAK-STAT signaling pathway, HIF-1 signaling pathway, etc.). Animal experiments also proved the therapeutic role of CAD in asthma by attenuating airway inflammation and mucus production and inhibiting the expression of Th1 and Th2 cytokines. Our study looked forward to providing a new perspective for developing traditional Chinese medicine on asthma therapy.
FundingThis work was funded by Zhejiang Provincia Natural Science Foundation of China (grant number LY15H270006).
Conflict of interestThe authors have no conflict of interest to declare.